- Learn how to build custom Python data connectors and further customize JMP’s Data Connector Framework with the Python Data Connector Demo, available now in the JMP Marketplace!
- See how to create experiments to support product design and ID useful product features. Register for June 12 webinar, 2pm US Eastern Time.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- formulations and their effects
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
formulations and their effects
Hi- I made 9 different chemical formulations using 6 ingredients (adding different wt% of each). I make a certain amount (g) of byproduct for each of the 9 formulations, the amount dependent upon how we formulate (ratios added). What would be the best way to analyze and make sense of the data? Ideally want to determine the best formulation to give the minimum amount of byproduct. I appreciate the help.
Tade
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
Tade,
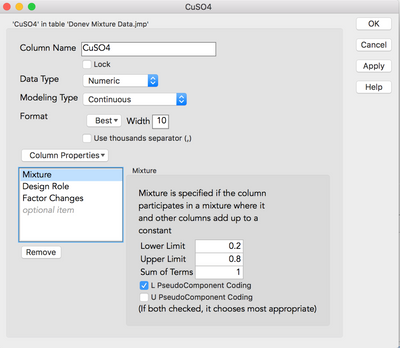
If you double-click your column what does your column information look like? I am curious to first ensure that you have the columns coded correctly. It should resemble this...
If it does resemble this then perhaps you do not have enough degrees of freedom to fit a Scheffe cubic. Have you just tried a main effects and two-way interactions (non-linear blends) model?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
I would recommend to:
1. Visualize your data, e.g. using the graph builder. Use graphs to give you an idea in which direction the data leads you. Check for potential outliers as well.
2. Then try to find an adequate model in the "Fit Model"-plattform. Make sure to set all factors in the model to mixture using the little red triangle next to "Attributes" in the fit model plattform. The JMP help gives an intro to the analysis of mixture experiments: A Chemical Mixture Example
Overall you might want to read about Design of Experiments for Formulations (Mixture Designs) for future studies if you are not already aware of that topic.
Hope that helps,
Sebastian
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
Building on what Sebastian has said, I think it would be most useful to you to set up a "dummy" mixture design of your experimental situation so you can see the resulting formulation design and the resulting design data table. Your weight percents can be entered as fractions that add to unity (1). If you examine the column attribute metadata from that "dummy" experiment you can then mimic the column attribute metadata in your actual formulation experiment JMP file and have the appropriate visualization tools like the mixture profiler and prediction profiler to evaluate and see the scripted model associated with a mixture design.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
Sebastian, Lou,
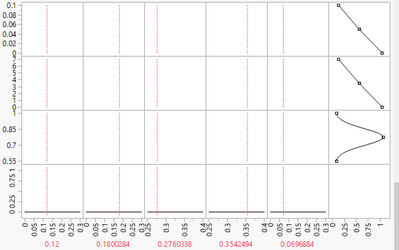
Thank you. I ended up using Scheffe polynomial in Mixture design. However, the Prediction Profile I am getting looks more like blank like this:
What am I missing? Thanks for your help.
Tade
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
It is impossible to know exactly why you see this without having the data. Best guess is that the range of Y-values shown on the profiler does not actually contain the responses. Try changing the Y-scale so that you can see the profile lines.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
Tade,
If you double-click your column what does your column information look like? I am curious to first ensure that you have the columns coded correctly. It should resemble this...
If it does resemble this then perhaps you do not have enough degrees of freedom to fit a Scheffe cubic. Have you just tried a main effects and two-way interactions (non-linear blends) model?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
Thank you both for quick responses.
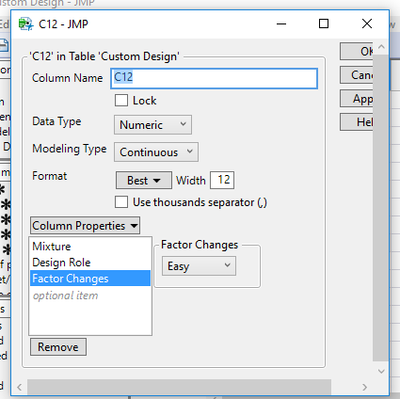
I tried changing the Y scale on the Prediction Profiler but didn't work. Here is my column information look like.
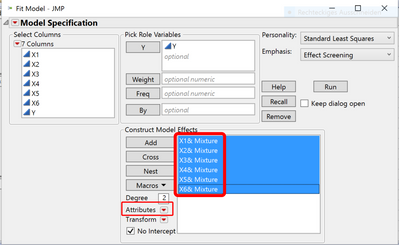
For the model, I selected Scheffe Cubic and it listed all the interactions. I have five factors and nine different combinations.
Thanks.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
I checked L PseudoComponent Coding for each column for factors.
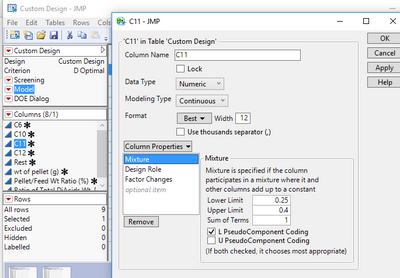
But I am getting the same results. No Y values no profile lines in the Prediction Profiler. Thanks for the help.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
I believe based on the number of experiments you are adding too many terms in the model. Please try to back off and just look at the main effects.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: formulations and their effects
Tade,
If you are using a Scheffe cubic with 5 mixture factors with 9 unique formulations then you have an over parameterized model, ie not enough df to estimate all terms. If you would be willing to sanitize your JMP dataset and share then I think the guidance you could receive may help you get to a solution faster. Also I am assuming those 9 runs did come from a properly designed mixture experiment which was generated to support a particular model. If that is not the case, then estimability of any particular arbitrary mixture model may or may not be possible depending on how those 9 points cover the mixture space. You have already received some great input from Lou and DanO so please do consider their feedback.
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us