- Learn how to build custom Python data connectors and further customize JMP’s Data Connector Framework with the Python Data Connector Demo, available now in the JMP Marketplace!
- See how to use Accelerated Life Testing (ALT) to evaluate reliability. Register for June 5 webinar, 2pm US Eastern Time.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- Fit Model- Repeated, Fixed and Random Effects
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Fit Model- Repeated, Fixed and Random Effects
Hi,
I am hoping for some guidance with a test I would like to run. We have a trial were birds were fed 2 diets (Control and Treatment). Every week (Day 7, 14, 21, 28) subjects were weighed to analyse performance data (weight gain, feed efficiency, mortality etc.). The exact same trial was repeated in the same facility two months later, using the exact same diets and trial protocol. I want to combine the results of both trials and analyse them together.
Please can somone advise how to do this? I am guessing I would need to add trial, treatment and day as fixed effects and do this via the Fit Model platform. Is that right? One of the trials performed better in general, but I am not interested in whcih trial performed better, only which treatment performed better overall. Should Day be added as a repeated effect or is there an option to do this? I am using JMP, but do not have JMP Pro, although I do have the repeated measures Add In, but not sure if this is what I should be using.
Also- what personality should I select, and can I add something as both a fixed and random effect?
Thanks in advance!
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Fit Model- Repeated, Fixed and Random Effects
Hi @Abbiegraham,
It sounds like you have a repeated measures experiment with one between-subject factor (diet) and one within-subject factor (time). You could consider the different runs of the entire experiment another between-subject factor, but if they really were identical I might ignore that fact since it will increase the complexity of the model.
You can run this model in standard JMP by using Analyze > Fit Model. Setting up the model effects will be easier using my add-in, which it sounds like you already have (https://community.jmp.com/t5/JMP-Add-Ins/Full-Factorial-Repeated-Measures-ANOVA-Add-In/ta-p/23904). I've attached a data set with the format the data need to be in.
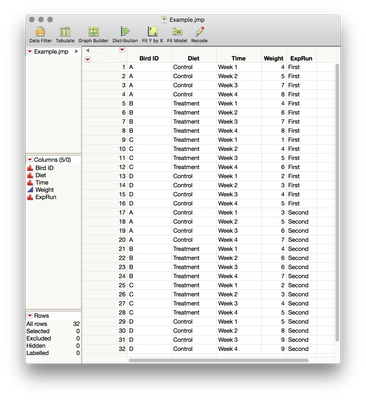
You'll see that I have included a column for the experimental run, but we won't be using it when we set up the model. If you were interested in seeing whether there is evidence for differences between the experiments you did a few months apart you could use that column as a between-subject effect (it's between-subjects in the same way that diet is between-subjects -- no single bird is measured at both levels of the factor).
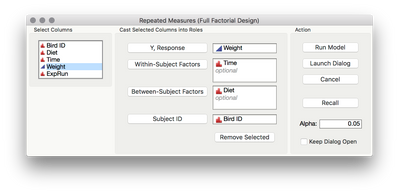
To set up the model using my add-in, we'll use Bird ID as the subject ID, Diet as a between-subject effect, time as a within-subject effect (and be sure that this column is marked as nominal), and weight as your Y.
This will return a linear mixed effects model equivalent to a standard repeated measures ANOVA.
I hope this helps!
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Fit Model- Repeated, Fixed and Random Effects
Hi,
I am a JMP Genomics user. My experiment is similar but each subject has thousands of genes to be tested with the row by row model.
So what do I do with genomics analysis in a similar experiment?
I do not get to fill in the "between" and "within" subjects factor. What would be the random or fixed factors? How do I decide on that?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Fit Model- Repeated, Fixed and Random Effects
Hi Melody,
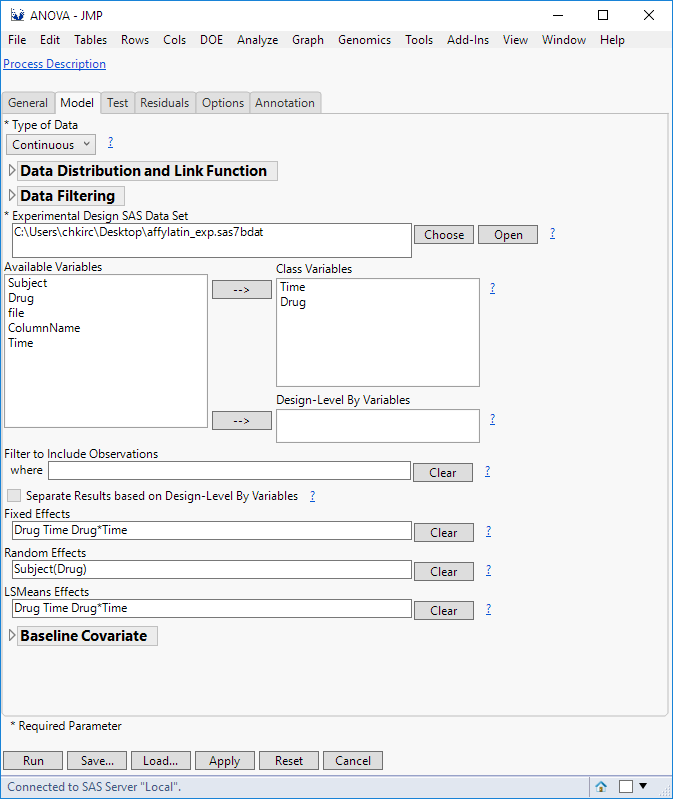
This can be done in JMP Genomics using the Fixed and Random effects seciton within the ANOVA dialog under Model tab (assuming that is what you are using). One is allowed to nest variables just like JMP is able to. Syntax is the similar, () instead of [], but one has to type them in instead of dragging and droping. Check out the below link and look at the Univariate Approach for the syntax of defining how the suject effect will be nested withing the between subject effect (drug). This would go into the random section, the rest are fixed effects and go into the Fixed Effects section.
http://www.jmp.com/support/notes/30/584.html
So in for this example, it would look something like this in JMP Genomics ANOVA dialog:
Here is another explaination of Nesting from a JMP point of view:
http://www.jmp.com/support/help/Launch_the_Fit_Model_Platform.shtml#65634
Hope that helps,
Chris
Data Scientist, Life Sciences - Global Technical Enablement
JMP Statistical Discovery, LLC. - Denver, CO
Tel: +1-919-531-9927 ▪ Mobile: +1-303-378-7419 ▪ E-mail: chris.kirchberg@jmp.com
www.jmp.com
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Fit Model- Repeated, Fixed and Random Effects
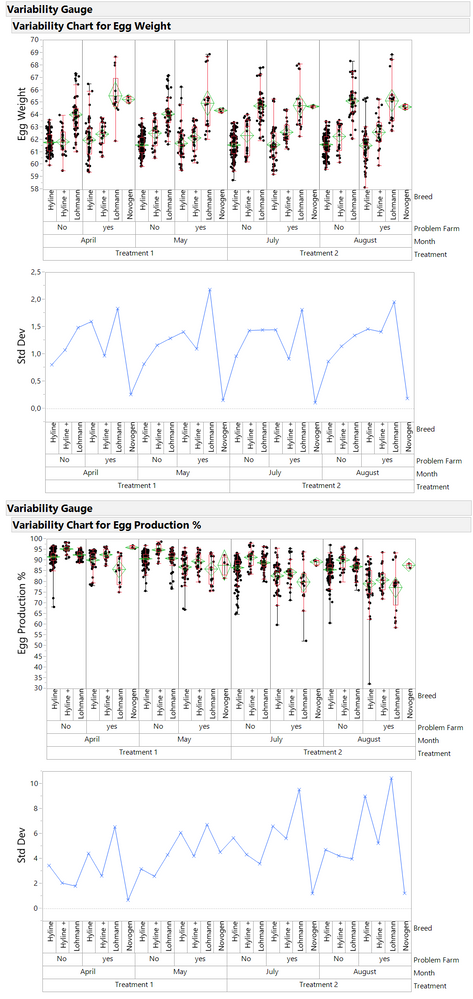
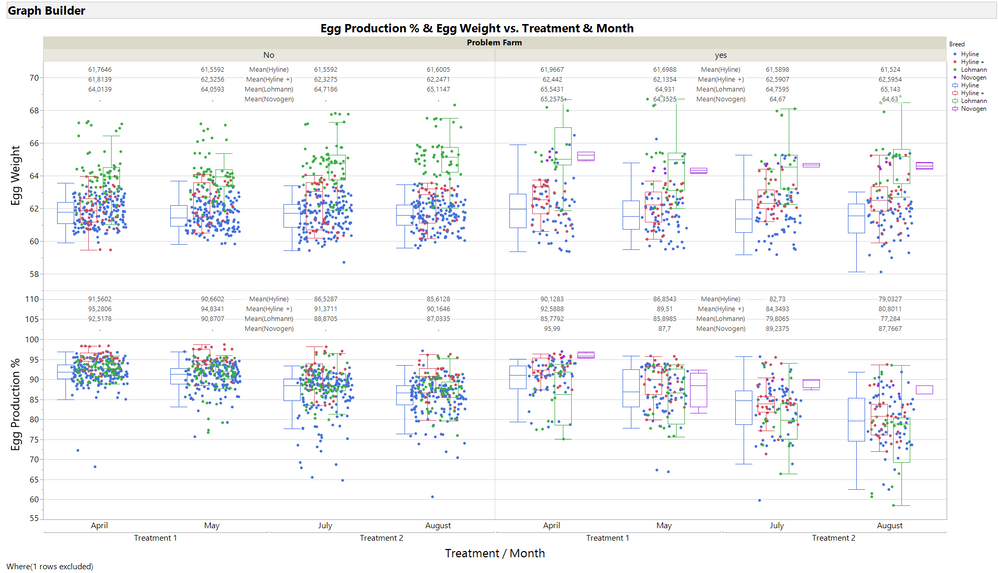
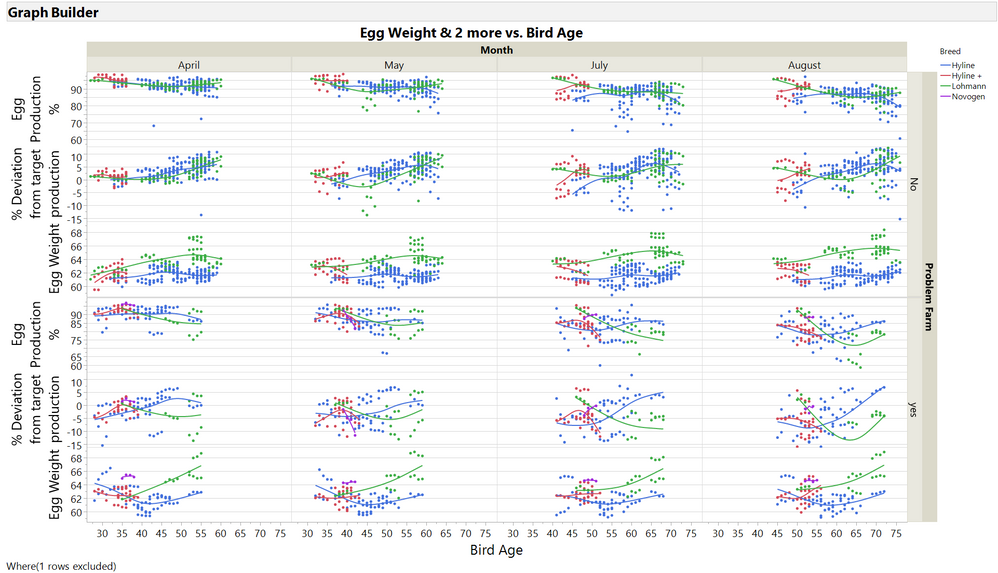
Was just checking different visualisation options.
Problem farm without information was set to "no".
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us