- See how to interactively organize and restructure data for analysis. Register for May 29 webinar, 2pm US ET.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- How can to remove the variation of a factor from data responses in a factorial d...
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
How can to remove the variation of a factor from data responses in a factorial design?
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Hi,
I've tried to attach your experiment as a .jmp table but had the same problem. I'll let the community manager know so it will hopefully be resolved soon.
I evaluated the design and it is almost completely orthogonal for main effects and 2-factor interactions (some slight correlation due to the excluded 'outlier' runs).
I haven't looked in detail at the approach that you took. I think it is probably okay but would not be recommended by a statistician.
My approach would be to fit a model including Genotype. I define it as a random effect. This is because my understanding is that it is a factor that you expect to affect the responses. But you don't actually care what the effect of Genotyope is. You just need to account for the effect so that it does not change your estimation of the effects about which you do care: Nutrient 1 and 2 and their interaction. Am I correct in that?
You should also define block as a random effect for the same reason.
So I fit a model with Block and Genotype as random effects and Nutrient1, Nutrient2, Nutrient1*Nutrient2 as fixed effects.This model tells you the effect of the nutrient factors having taken into account the variation in the responses due to Block and Genotype.
If you try this do the results broadly agree with your analysis? It looks like Nutrient 1 has a significant effect on Variable 1 and the other the effects are not significant.
Regards,
Phil
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Yes, that is possible. From any model in JMP you can save residuals, e.g. from Fit Y by X: https://community.jmp.com/t5/Discussions/How-to-save-residuals-in-Fit-Y-by-X-platform-for-two-contin.... You can also save residuals from Fit Model and other modelling platforms.
Is it correct? Well there is no right or wrong really. You need to think about what you are doing here. You also need to consider how good your data is for separately estimating the effect of A. Are the effects of A, B and C orthogonal in your experimental design? See my recent blog post for an understanding of what I mean.
I would say that it is more common to include A as well as B and C in a multiple regression model. That way the variation in biomass due to A is captured by the model and will allow you to understand the effect of B and C.
Hope that helps.
Phil
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Dear Phil,
Thank you so much for your time and clear explanation.
I saw the links that you mentioned. I think the effects of A*B and A*C and B*C on variables in my experiment look like to design 2, so they are not orthogonal.
Could you please check out the process that I did to remove the effect of A from my variables?
1. I saved the residuals from A with each variable.
2. I run full factorial (B and C) on saved residuals. Then I saved the residuals as a new variable.
Now, the effect of A has been removed from the new variables.
Do you think it is correct?
Thank you so much for your time,
Cheers,
Milad
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Hi,
It is difficult to answer from your description. I am not sure about the approach that you have taken. Are you able to share the data or an anonymised version? Or can you illustrate what you are trying to do with one of the JMP sample data sets? Maybe the "2x3x4" example is similar:
Open("$SAMPLE_DATA/Design Experiment/2x3x4 Factorial.jmp")Regards,
Phil
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Dear Phil,
Thank you so much for your time to advise me.
I have attached the factors with a part of my data. Please find it. I couldn't attach the .jmp file, I saw this message: "The contents of the attachment doesn't match its file type.". So I put the image of steps and attached the excel sheet.
This experiment is factorial under RCBD. Genotype (4 levels, A, B, C, and D), Nutrient 1 (2 levels, 10 and 40), and Nutrient 2 (2 levels, 25 and 100) are factors in 4 blocks. Also, "Variable 1" and "Variable 2" are variables that I measured during the experiment period. In this file, I transformed the variables in logarithm.
For example, different "Genotype"s had a different effect on "Variable 2". While I just need the effects of "Nutrient 1" and "Nutrient 2" in the model. So, I want to remove the effect of "Genotype" from "Variable 1" and "Variable 2". For this purpose, first I used "Fit Y by X" for each variable by "Genotype", then I saved the residuals for each variable.

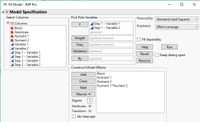
In the second step, I used these residuals in "Fit model" by "Block, Nutrient 1, Nutrient 2, Nutrient 1*Nutrient 2" as full factorial. After that, I saved the residuals that I obtained from this method and used them as new variables.
I mentioned, "Step 1" and "Step 2" in "Data table" in this file. Also, "Step () - Variable ()" are residuals that I saved from each step for each variable. (I couldn't attach .jmp file). For these variables, I just removed two outliers that you can see them in this file (row 20 and 61).
Thank you so much,
Sincerely yours,
Milad
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Hi,
I've tried to attach your experiment as a .jmp table but had the same problem. I'll let the community manager know so it will hopefully be resolved soon.
I evaluated the design and it is almost completely orthogonal for main effects and 2-factor interactions (some slight correlation due to the excluded 'outlier' runs).
I haven't looked in detail at the approach that you took. I think it is probably okay but would not be recommended by a statistician.
My approach would be to fit a model including Genotype. I define it as a random effect. This is because my understanding is that it is a factor that you expect to affect the responses. But you don't actually care what the effect of Genotyope is. You just need to account for the effect so that it does not change your estimation of the effects about which you do care: Nutrient 1 and 2 and their interaction. Am I correct in that?
You should also define block as a random effect for the same reason.
So I fit a model with Block and Genotype as random effects and Nutrient1, Nutrient2, Nutrient1*Nutrient2 as fixed effects.This model tells you the effect of the nutrient factors having taken into account the variation in the responses due to Block and Genotype.
If you try this do the results broadly agree with your analysis? It looks like Nutrient 1 has a significant effect on Variable 1 and the other the effects are not significant.
Regards,
Phil
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Dear Phil,
Thank you so much for your time, it was great.
Yes, the effect of "Genotype" is not important for me. I want to remove it from my data. I think maybe I can evaluate variables in a better condition. I think your solution is great.
Do you think that I can use these residuals as new variables to reduce the "Genotype" effect? Because I want to use them in the structural equation model.
Best regards,
Milad
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
I'm not sure what residuals you are talking about now. You would not want to use the residuals from the model that I have suggested because those residuals would only tell you about the variation that is not explained by Block, Genotype and Variables 1 and 2.
Structural Equation Modelling is quite a difficult methodology to understand (and is not possible in JMP currently). I am not sure how you would apply SEM here but I suggest that you make sure that you are comfortable in your understanding of regression modelling first.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Dear Phil,
Thank you for your message.
I mean the residuals that I saved them from the model that I ran by "Genotype" and "Block" randomly.
I believe that I should work more on it until I use them.
I want to run a sem with R, I just want to calculate and prepare data in JMP.
Thank you so much,
Milad
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How can to remove the variation of a factor from data responses in a factorial design?
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us