- New to JMP? Let the Data Analysis Director guide you through selecting an analysis task, an analysis goal, and a data type. Available now in the JMP Marketplace!
- See how to install JMP Marketplace extensions to customize and enhance JMP.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- Re: Windows 11 > JMP 19 > Fit Model > Standard Least Square > How to handle unus...
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Windows 11 > JMP 19 > Fit Model > Standard Least Square > How to handle unusual data distribution?
Hi JMP Community,
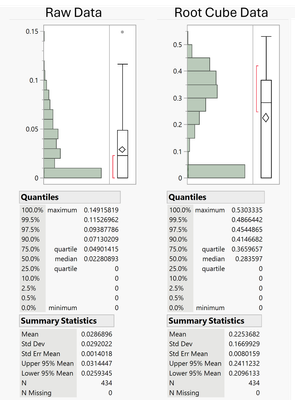
I am working with cell counts derived from tissue biopsies that exhibit an unusual distribution: some biopsies have no cells (zero counts), while others have low percentages (see example below). A simple Root Cube transformation yields a decent distribution of non-zero counts, but the overall distribution of the transformed data remains zero-biased.
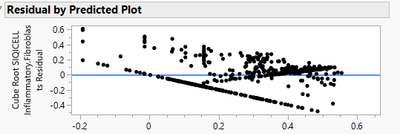
As expected, if I use this data in a Standard Least Squares Fit Model, the Residuals are not normally distributed (see below)
What would be your recommendation to test if these cell counts are associated with multiple covariates, including interactions?
Notes:
- The absence of cells (zero counts) is scientifically meaningful.
- I cannot easily share the actual data due to sensitivity.
Thank you.
Best regards,
TS
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Windows 11 > JMP 19 > Fit Model > Standard Least Square > How to handle unusual data distribution?
1. Determine when result is 0 or different from 0 (binomial distribution)
2. For the cases when results are different from 0, fitting a standard least squares model. You can still apply a transformation on your raw data (if needed !) like Box-cox transformation.
Best,
"It is not unusual for a well-designed experiment to analyze itself" (Box, Hunter and Hunter)
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Windows 11 > JMP 19 > Fit Model > Standard Least Square > How to handle unusual data distribution?
Hi @Thierry_S,
Do you have JMP Pro ? Using Generalized Regression models, you can specify one of the zero-inflated distributions.
I think zero-inflated Poisson distribution could work on your count raw data. If you're using JMP, maybe you could Split your modeling in two parts:
- Determine when result is 0 or different from 0 (binomial distribution)
- For the cases when results are different from 0, fitting a standard least squares model. You can still apply a transformation on your raw data (if needed !) like Box-cox transformation.
Hope this suggestion may help you,
"It is not unusual for a well-designed experiment to analyze itself" (Box, Hunter and Hunter)
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Windows 11 > JMP 19 > Fit Model > Standard Least Square > How to handle unusual data distribution?
Dear Victor,
Thank you for helping me with this problem. Unfortunately, I do not have access to JMP Pro at this time. Still, I have access to the Generalized Linear Model in JMP 19 "Basic", but the Poisson Distribution does not seem to fit all the data. Indeed, when the cell type is more abundant, the non-zero sub-population distribution tends to approximate a normal distribution.
I wonder if binning the data as an ordinal variable could help here?
Best,
TS
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Windows 11 > JMP 19 > Fit Model > Standard Least Square > How to handle unusual data distribution?
1. Determine when result is 0 or different from 0 (binomial distribution)
2. For the cases when results are different from 0, fitting a standard least squares model. You can still apply a transformation on your raw data (if needed !) like Box-cox transformation.
Best,
"It is not unusual for a well-designed experiment to analyze itself" (Box, Hunter and Hunter)
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Windows 11 > JMP 19 > Fit Model > Standard Least Square > How to handle unusual data distribution?
Without SME, it is difficult to answer. First, what questions are you trying to answer? Second, what is the data source? Third, how confident are you with the measurement system? Are there other measure you can take? How confident are you the process for taking the biopsies is consistent? What is the model you are trying to fit? Consider if you split the data (as suggested by Victor), the question you must ask is are there different factors responsible for effecting cells counts as the factors effecting no cells? If so, it would make sense to have the two Y's (2 categories, binomial) and continuous cell counts for the data.
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us