- Learn how to build custom Python data connectors and further customize JMP’s Data Connector Framework with the Python Data Connector Demo, available now in the JMP Marketplace!
- See how to use Accelerated Life Testing (ALT) to evaluate reliability. Register for June 5 webinar, 2pm US Eastern Time.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- Prediction Profiler
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Prediction Profiler
Hi all,
I'm trying to fit a model to full factorial experiment (4 factors, curvature likely in all).
I have a tight data fit (RSq=0.81). However, the prediction profiler is not generating any graphs. We didn't add centerpoints across all factors in the first pass of the experiment. Two of the factors we have center points for. If I simplify the model for screening and treat the data as continuous, I'm getting a fit, but it's not accurate. I also added in the centerpoints we have as "discrete numeric" and then performed a fit model. This fit is better, but again, it's not perfectly aligned with the data. Is this JMP's way of telling me it's lacking all of the data to make accurate predictions? We will be running another pass with centerpoints, I'm more asking if this is JMP's way of telling me it's lacking data.
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Prediction Profiler
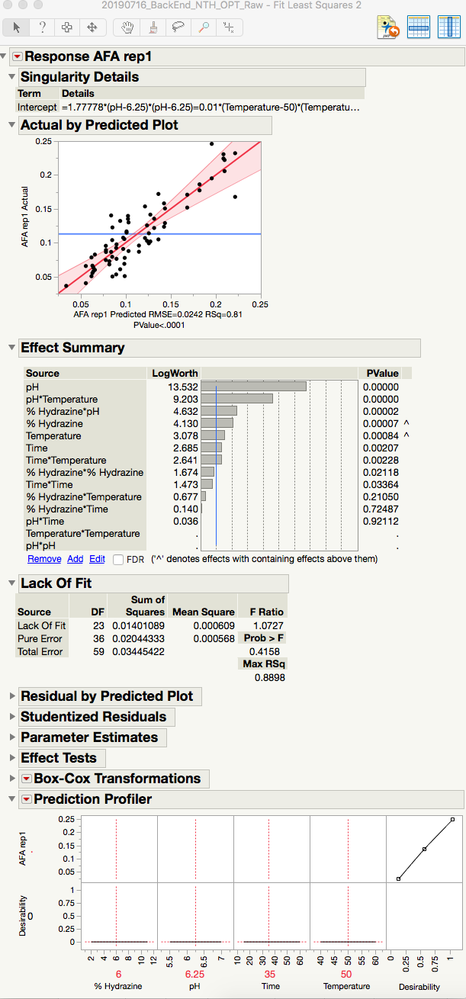
Do you see the Singularity Details report at the top of the Fit Least Squares platform? You data does not support the estimation of this model. In particular, the pH squared and Temperature squared terms must be removed. You can easily select and remove them from the Effect Summary near the top.
What do you get now?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Prediction Profiler
Do you see the Singularity Details report at the top of the Fit Least Squares platform? You data does not support the estimation of this model. In particular, the pH squared and Temperature squared terms must be removed. You can easily select and remove them from the Effect Summary near the top.
What do you get now?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Prediction Profiler
Oh, that's great! And JMP 14 is way more user friendly than the JMP 10 I was used to. Your solution make sense as those two factors are those that I do not have centerpoints for.
Thanks a lot Mark!
Jack
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Prediction Profiler
If you want to add observations, then you should use Augment Design. Generally, you design the initial experiment with Custom Design. If you determine that more data is necessary for any reason, Augment Design uses the initial experiment and adds the best runs to it.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Prediction Profiler
You appear to have a large number of runs for only four factors. Your initial experiment has two two-level factors and two three-level factors. That means a full factorial design with 2 x 2 x 3 x 3 = 36 runs but you have 59 degrees of freedom in the sum of squares for the error alone. The model is like to contain the intercept (1), main effects (4), two-factor interactions (6), and squared terms (4) or 15 terms in all.
What design method did you use?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Prediction Profiler
There are 72 data points because we ran two replicates of the full factorial. I’m basically back-ending data into a model. We didn’t use JMP to design the experiment. Normally, I do use the software to design and will do so going forward.
Thanks for the help.
Jack
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us