- Learn how to build custom Python data connectors and further customize JMP’s Data Connector Framework with the Python Data Connector Demo, available now in the JMP Marketplace!
- See how to use Accelerated Life Testing (ALT) to evaluate reliability. Register for June 5 webinar, 2pm US Eastern Time.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- Paired t test
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Paired t test
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
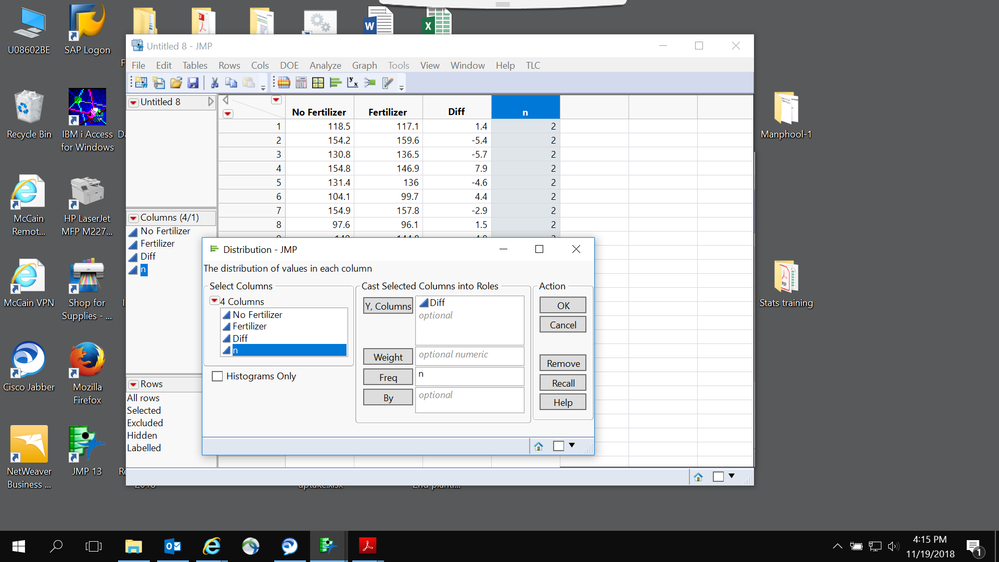
No, do not add a frequency column. Each pair is 1 data point in a paired t-test. A paired t-test is mathematically equivalent to a 1-sample t-test on the paired differences.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
Thanks so much for your quick reply. I understand that the paired t test can be performed by using the following four methods: I am wondering whether all four methods are equally good. If not which one is the best and why? I got identical results.
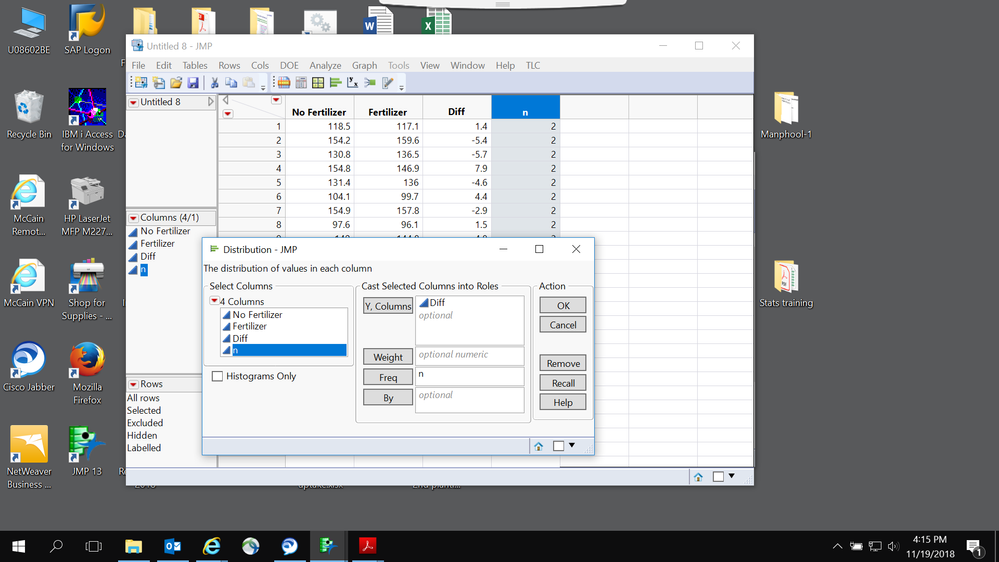
1. Analyze > distribution- mean test
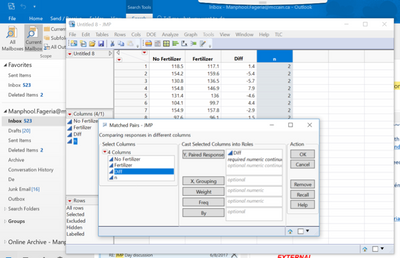
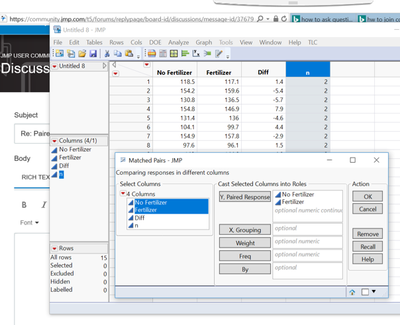
2. Analyze-specialized modeling-matched pairs
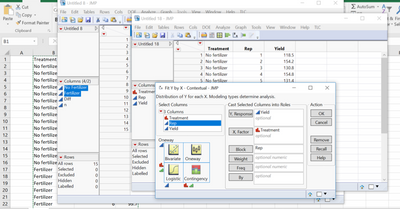
3. Fit Y by X > compare means> Each pair, students t test- detailed comparison report
4. Fit Model platform to do a two-way ANOVA
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
In #4 you should have Rep as a random effect in order to get an equivalent result to the other methods. There is actually another way, which is to plot your Diff column in the Distribution platform and do a 1-sample t-test. Since they should all give the same answer, you could say they are equally good. The Matched Pairs platform is specialized for this problem, so you'll get the most specialized/relevant output in your report there. However, that platform requires the data to be in a different shape than the usual 'long' format. If I'm want to avoid splitting my table to get it in the right shape, I might just use Fit Y by X with the pair ID column as the block, or Fit Model with a random effect on the pair ID column.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
Thanks so much again,
Sorry, could you please expain the following:
1. How to use rep as a random effect in #4? Could you please explain it by including a table.
2. Could you include another table explaining Fit Y by X with the pair ID column as the block.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
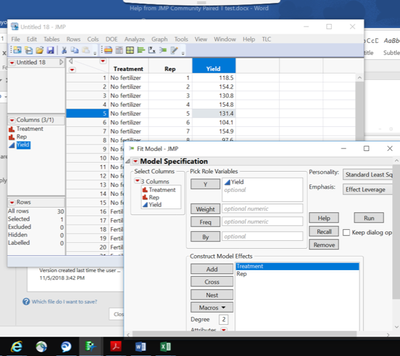
Here's an example using Gosset's Corn data from the JMP's Sample Data Library. I added a pair ID column and then stacked the 2 yield columns (Kiln and Regular) to get to a long data format. Make sure the pair ID column has a nominal modeling type.
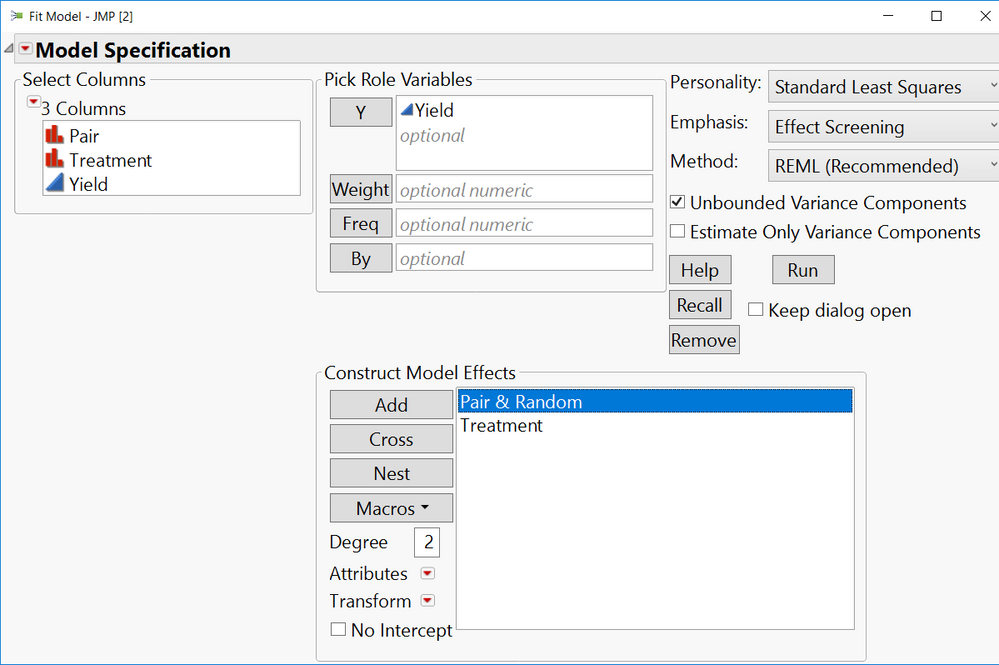
From Fit Model:
1. Add the Yield column as the Y, Response variable.
2. Add the Treatment and Pair columns into the Construct Model Effects box.
3. Select (highlight) 'Pair' in the Construct Model Effects box and make it random by clicking the Attributes red-arrow > Random.
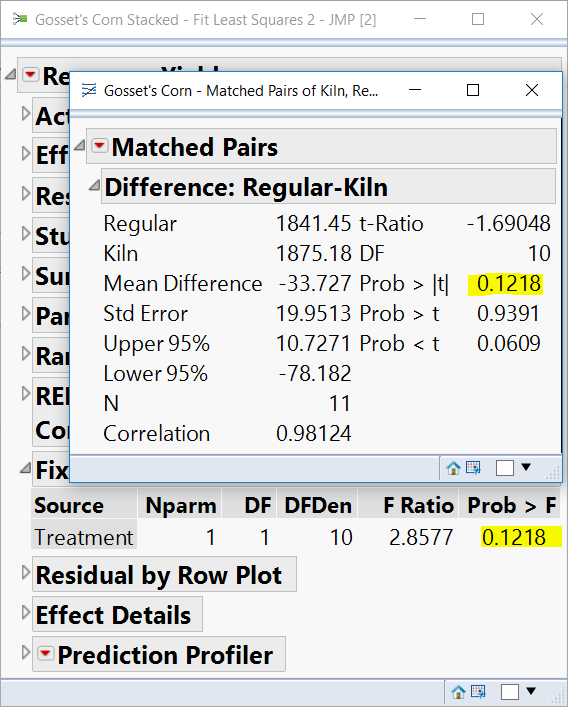
You'll see the p-value for Yield has the same p-value as the Matched Pairs platform for the same data in the original wider format.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
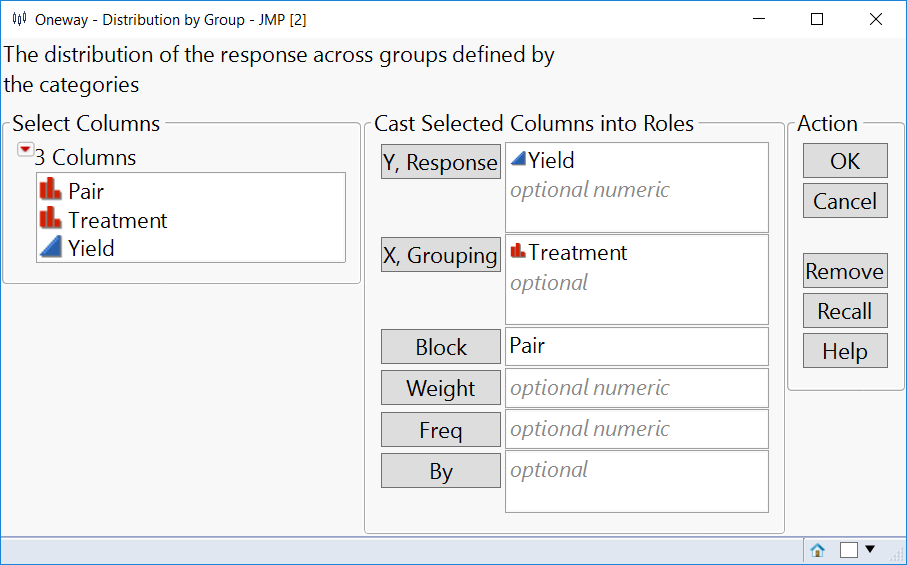
Here's how to get the same p-value in Fit Y by X with the same stacked data table for Gosset's Corn:
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
Please clarify the basis for pairing the samples. How were two samples paired together?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
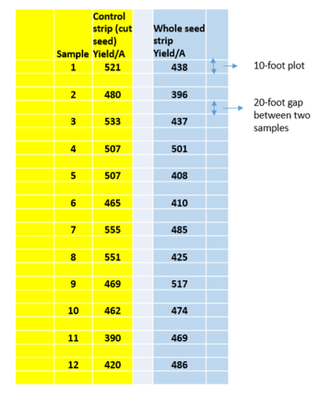

Hi Mark,
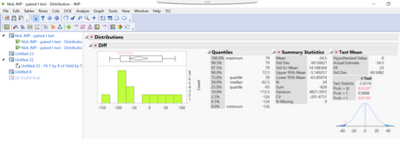
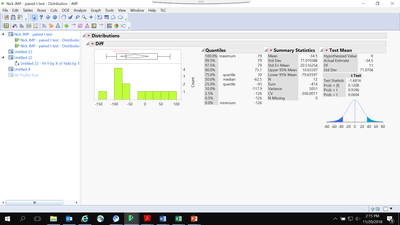
We wanted to compare between two seed types in potatoes (planting Cut seed vs planting whole seed). Cut seed is used as control which is a common practice. This trial was conducted at the grower’s field. The field was divided into two strips blocks, the first block (in yellow) was planted with control (cut seed) and the second block (light blue) was planted with the whole seed. These strip blocks are really long 400-foot. In total, twenty four samples (12 from each strip block) were harvested. Each sample was harvested from a 10-foot long row plot and converted the yield in CWT per acre. The field variation was huge. The trial was conducted at grower’s field. So, we could not randomize the treatments. I have analyzed the data using the paired t with distribution and test mean, see below:
My question:
I understand that we should use the paired t test.
One of the great statistician suggested that we should create an extra column frequency and use it for the analysis. I have analyzed the data with and without using the frequency.
I wanted to know which method is incorrect and why?
Which method is correct?
I would appreciate your time.
Best regards
- I have created difference and frequency columns and both
See the results below: They are significant P >0.0232
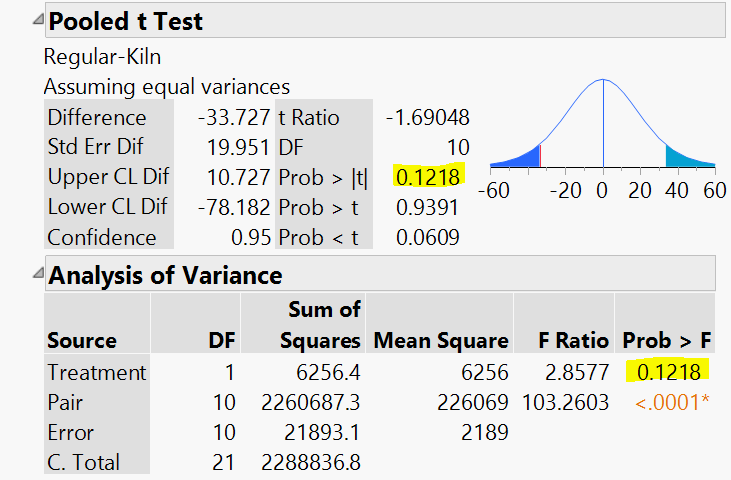
- I have created difference and frequency column but this time I did not use the frequency column
Results see below: Non significant P >0.1208
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Paired t test
You description of blocks is yellow and blue, so the blocking (pairs) does not make sense,
If the cut and whole seed plantings occurred next to each other, but the 12 strips are separated over a variable fied, the blocking (pairs) make sense. Is the field homogeneous within a pair? That assumption is important.
Yu do not need any more than the two response columns, CONTROL and WHOLE SEED. Use the Matched Pairs platform in the Analyze menu. Enter both of the data columns in the Y role in the order above. Do not enter any other data columns in this analysis.
It seems that you expect a higher yield for the whole seed treatment (alternative hypothesis), so an upper-tailed paired t-test is called for.
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us