Here are three ways to create clustered bar charts using JMP. I discussed this in Chapter 5 of my book Biostatistics Using JMP, and one of the ways is very similar to a recent example shown by @XanGregg. However, two extensions of the basic clustered bar chart might be important depending on the story you are telling, and that's what we're covering here.
The three approaches we'll look at here are: 1) clustering by group (as shown in the recent example), 2) clustering by variable, and 3) clustered bar charts with data points. We'll also add error bars for good measure in the first two bar charts; due to clutter issues, the third bar chart will not show the error bars. We'll use the Fisher Iris data for this example, which you can load by going to Help > Sample Data > See an Alphabetical List of all Sample Data Files and selecting the Iris file.
Figuring out how to do all of this in JMP came up during some real-world work, and it involved remembering how to organize data effectively. @Bill_Worley was able to help put the finishing touches on what I was missing (everyone needs help from the masters sometime). Needless to say, the process in JMP is much easier and quicker to achieve than it was in MATLAB.
Clustering by Group
Clustering by group is the simplest method and was presented in Xan Gregg's example. Here, we consider the data table as is. We can cluster by group via Graph Builder or Charts, but we'll focus on Graph Builder since it is very versitile and facilitates the next approach.
In Graph Builder, drag the four data columns (Sepal length, Sepal width, Petal length, and Petal width) to Y. Next drag the group (Species) to X. Click on the bar chart icon  . We'll also add confidence intervals through the error bar drop-down menu. Once we're happy with the look, we can then click Done. We now see a result just like that shown below and as shown previously.
. We'll also add confidence intervals through the error bar drop-down menu. Once we're happy with the look, we can then click Done. We now see a result just like that shown below and as shown previously.
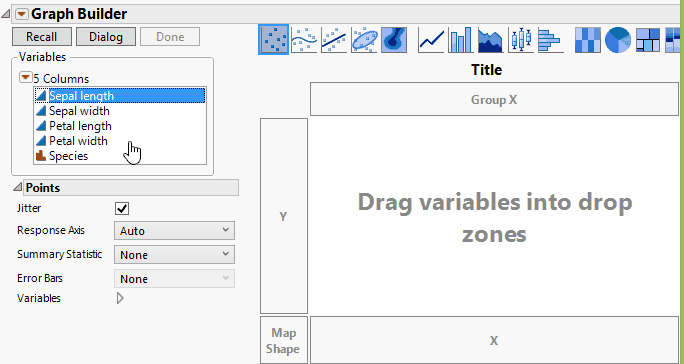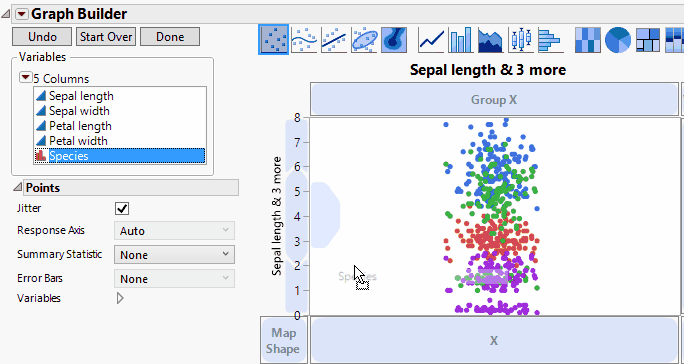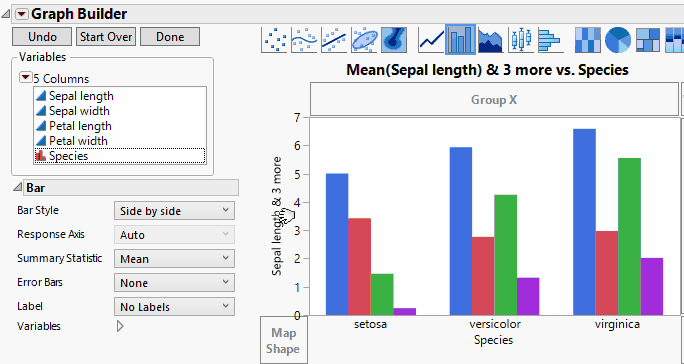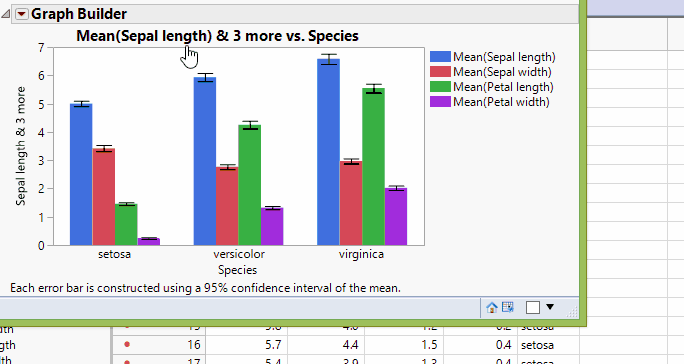 Fisher Iris Variables, Grouped By Species
Fisher Iris Variables, Grouped By Species
However, what if you're interested in a variable's mean response differs by group? For instance, what if we wanted to see how sepal length differed across species? It would be very difficult using the above chart. So, we'll have to cluster by the variables to much such comparisons.
Clustering by Variable
Unfortunately, the data isn't set up to immediately cluster by variables. In order to do this, we need a data variable that indicates the group (Species in this case) and a data variable that indicates that variable name. We have a few options to do this. We could try Tabulate, but we would not be able to add error bars using its result. So we'll create a new data table using the Stack function. We'll stack all of the columns of data, but not the Species.
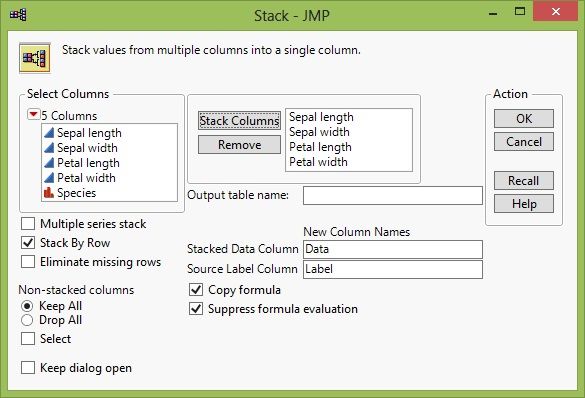 Stacking Data
Stacking Data
Now we have a data table that has three columns: Species, Label (the data variable names), and the data itself. The data table is now 600 rows long since Fisher Iris has three species, with 50 observations each of the four variables.
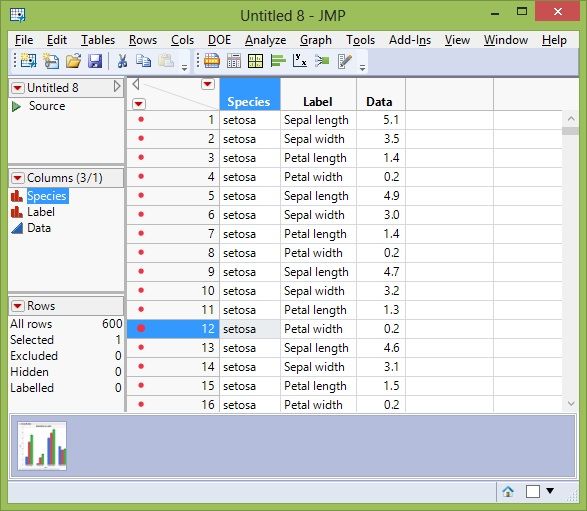 Stacked Data Table
Stacked Data Table
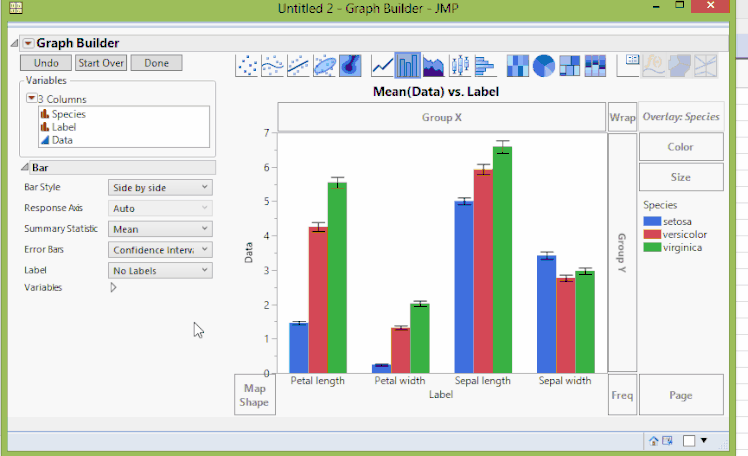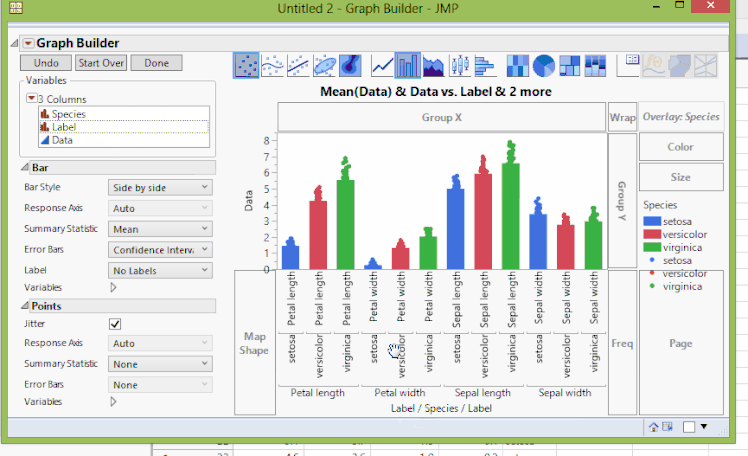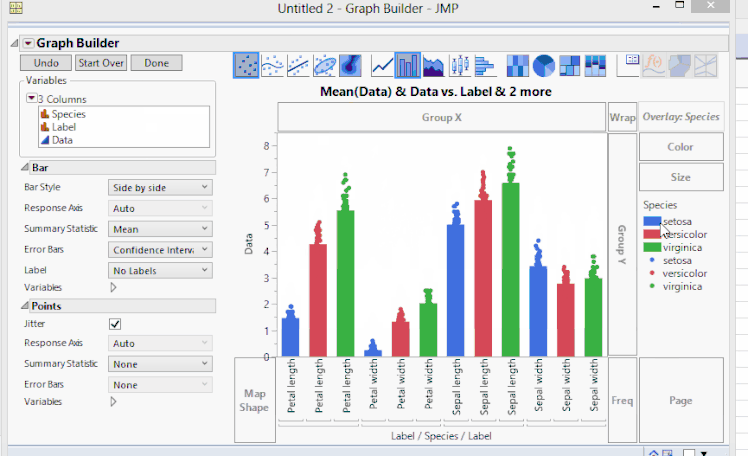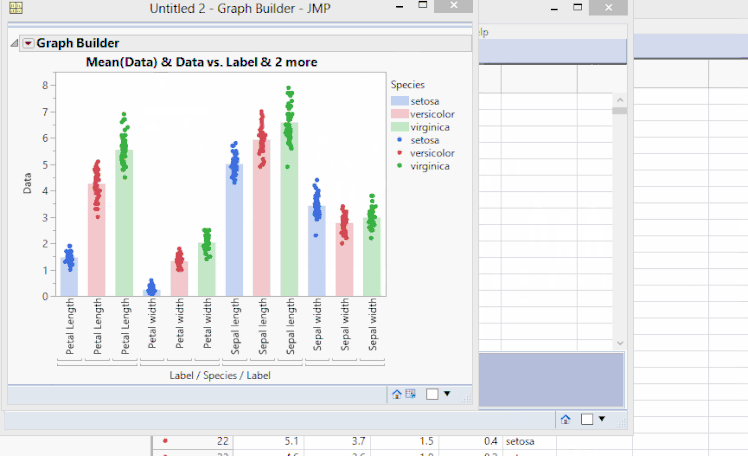
To create the graph we've been looking for, we'll go back to Graph Builder, and now we'll drag data to Y, Label (which contains the original variable names) to X, and Species to Overlay. We'll again add confidence intervals through the error bar drop-down menu. We can then click Done once we're happy with the chart. The result now shows the same data as before, but with a representation that might be more useful in some situations.
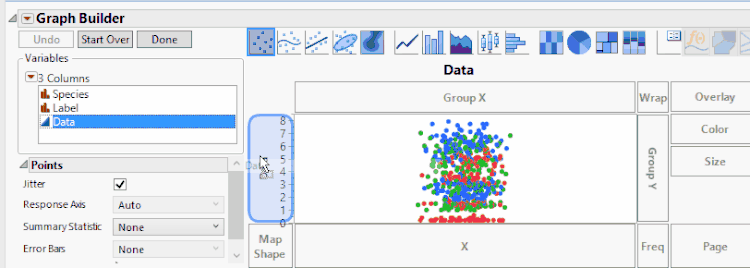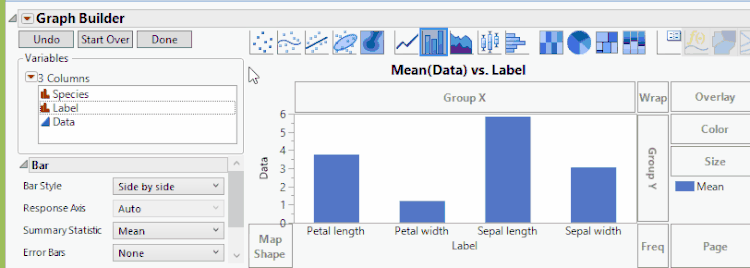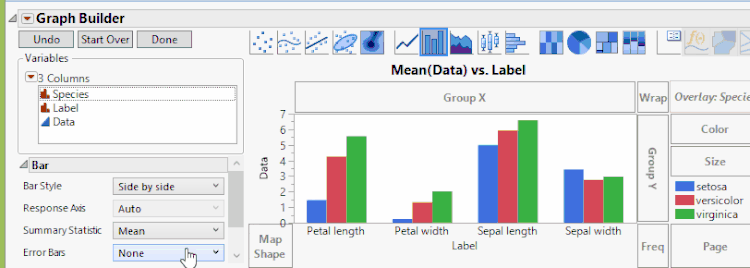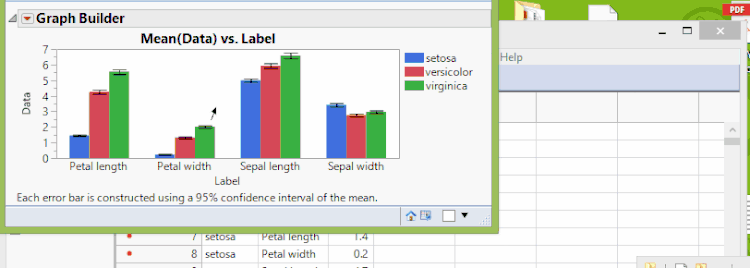 Fisher Iris Variables, Grouped By Variable
Fisher Iris Variables, Grouped By Variable
Clustered Bar Charts with Data Points
For full disclosure of data ranges, it might be beneficial to show the actual data points. However, we won't be able to show the error bars at the same time; it would look too cluttered.
To create a bar chart where we have points overlaying the bars, we'll continue with the clustering by variables example. In Graph Builder, we'll have to turn on both bar charts and points. Since bars are already on, we'll have to turn on the points. We can do this by holding the Shift key and pressing the points button. Now we see that the points all line up on only one of the bars. We can tell JMP which points go to which bars by dragging the Species to X. However, now the grouping is not what we had before. We can fix this by dragging Label again to X. Mostly, this graph is what we want to see.
But, the X labels are rather cluttered since we have three nested levels to group by Species and Label. We can fix this by changing Axis Settings and unclicking Show Tick Labels on the superflous labels. For one last finishing touch, we can make the bars slightly transparent to show the points more effectively. We can do this by right-clicking on each bar's entry in the legend and changing the Transparency value (from 1.0 to 0.3 in this example). The result now shows the same data as before, but with yet another representation that might be more useful.
 Showing Original Data Points when Clustering by Variables
Showing Original Data Points when Clustering by Variables