- JMP User Community
- :
- File Exchange
- :
- JMP Add-Ins
- :
- XGBoost Add-In for JMP Pro
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
New for JMP 17 Pro: An Autotune option makes it even easier to automatically tune hyperparameters using a Fast Flexible Filling design.
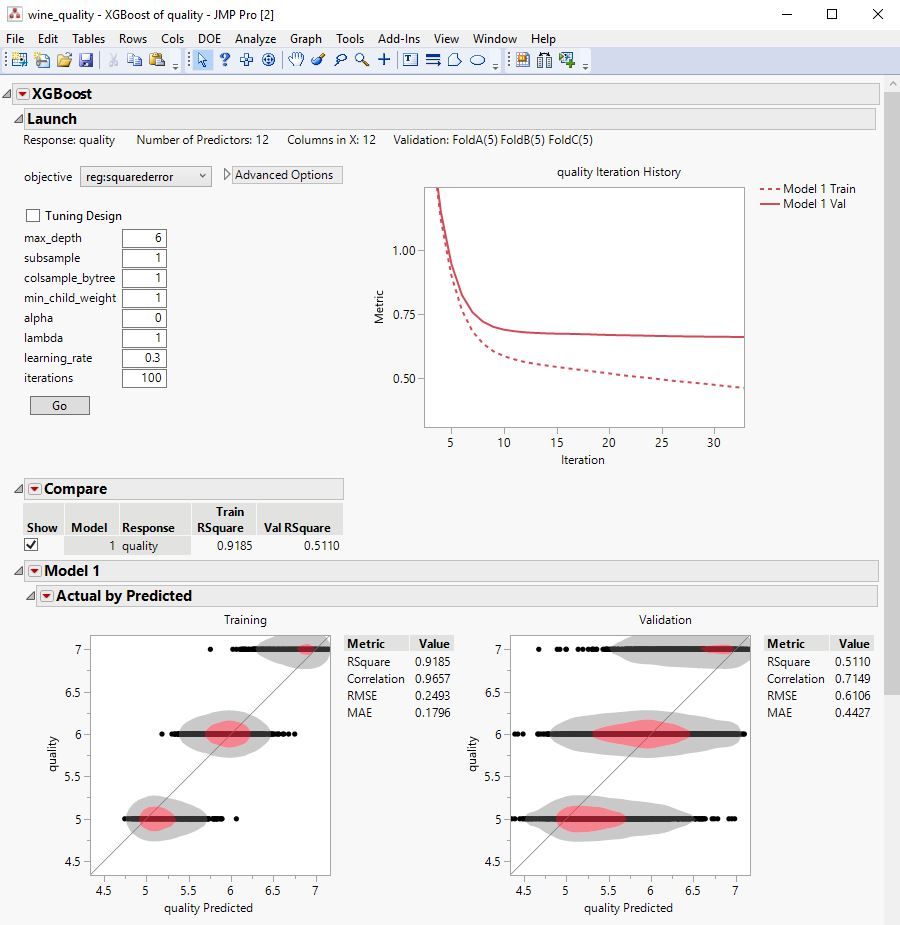
The XGBoost add-in for JMP Pro provides a point-and-click interface to the popular XGBoost open-source library for predictive modeling with extreme gradient boosted trees. Value-added functionality of this add-in includes:
• Repeated k-fold cross validation with out-of-fold predictions, plus a separate routine to create optimized k-fold validation columns, optionally stratified or grouped.
• Ability to fit multiple Y responses in one run.
• Automated parameter search via JMP Design of Experiments (DOE) Fast Flexible Filling Design
• Interactive graphical and statistical outputs
• Model comparison interface
• Profiling Export of JMP Scripting Language (JSL) and Python code for reproducibility
Click the link above to download the add-in, then drag onto JMP Pro 15 or higher to install. The most recent features and improvements are available with the most recent version JMP Pro, including early adopter versions.
See the attached .pdf for details and examples, which can be found in the attached .zip file (along with a journal and other examples used in this tutorial).
Related Materials
Video Tutorial with Downloadable Journal: XGBoost Add-In for JMP Pro (2020-US-45MP-540)
Add-In for JMP Genomics:
Add-In from @Franck_R : Machine Learning with XGBoost in a JMP Addin (JMP app + python)
Congrats Russ, this addin looks indeed very promising and useful!
Thank you Franck, dittos for yours! On Kaggle it appears LightGBM is the most popular so that may be a direction we would want to pursue down the road, and also CatBoost.
Hello Russ
you are writing in the documentation that XGBoost has an "efficient handling of missing data". But you also marked it with a "?". Please, could you explain how the add-in deals with missing data?
Thanks in advance
Markus
@markus Thanks for the question on missing values; a few others have raised it as well. For Nominal predictors, the add-in builds 0-1 one-hot / dummy columns and treats missing values as a separate level. For Continuous predictors, the add-in calls the XGBoost executable directly and feeds the missing values. When splitting a tree node, the algorithm assigns the missing values to the split that best improves the fitting criterion. Suggest web searching for more details. e.g. I was just looking at https://towardsdatascience.com/xgboost-is-not-black-magic-56ca013144b4 and it provides some nice commentary and some further research.
I've found this dual approach to work well across a wide variety of applications. It sure is nice not to have to fiddle with and impute missing values before modeling. Do be careful though if missingness is at least moderately predictive of the response, as it may signal data deficiencies that should be addressed.
Analyze > Screening > Explore Missing Values is a great starting place for discovering missing data structure.
Thanks Russ!
This addin is awesome. The bonus addin to create a k-fold validation column is very nice! I love the fact that all other "standard" features are already there, including the profiler and columns importance.
Hey JMP! You should allow using a k-fold validation column for all predictive platforms, just like what is done here!!!
Hi XGBoosters, There is a known crash in JMP Pro 16.0 on Windows that has been fixed in 16.1. Apologies for any inconvenience and please feel free to provide feedback any time.
Hello Russ
I am using JMP Pro15.21 in Win 7 64-bit OS.
I tried installing XGBoost add.
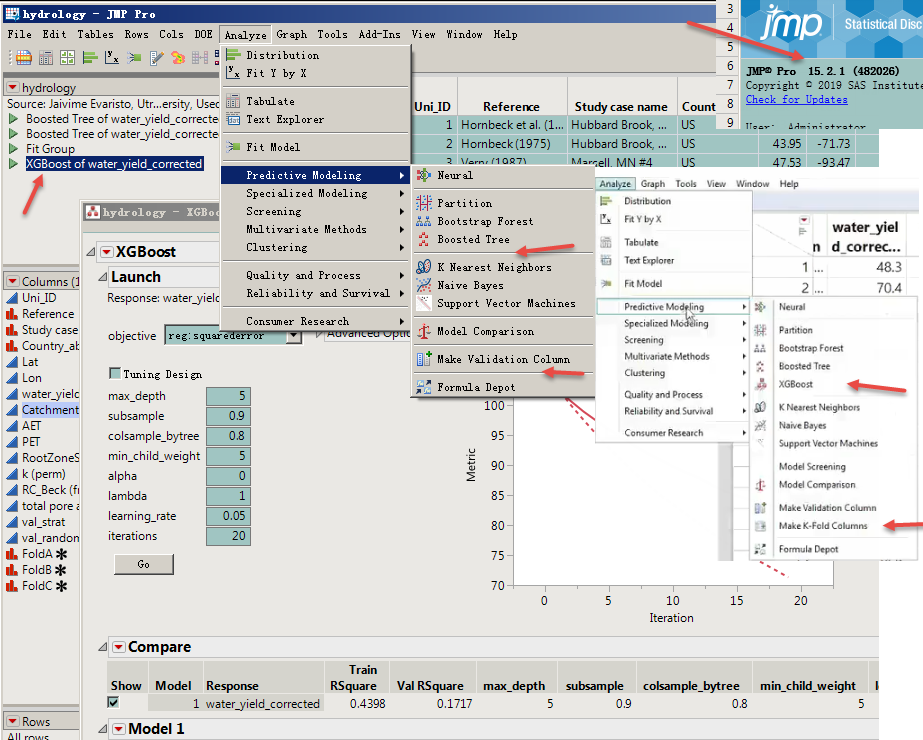
Cannot find options like those in the video in the analysis menu.
But open the "hydrology.jmp" file and can run the "XGBoost of water_yield_corrected"JSL inside.
I wonder why? Thank Russ!
Hello @lwx228 Apologies for the delay in replying. Look under your Add-Ins menu for the routines, and use View > Customize > Menus and Toolbars if you want to move them.
Hello Russ,
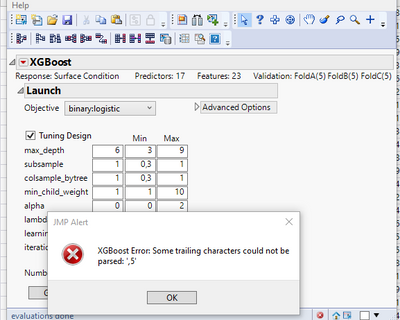
since longer I like to use the XG Boost again. In the meantime I have updated to JMP-Pro 16. I realize several error messages. The first ones I could overcome by re-installing the Add-in. Creating the 3 validation columns work fine. But running the Addin produces always this error:
"Some trailing characters could not be parsed: ',5'".
What is wrong and what should I do?
Thanks for help in advance!
Markus
Hello @markus , If you can send an example to russ.wolfinger@jmp.com I can investigate. Looks like it has something to do with ',5' being in column names or values and is perhaps related to non-English setting. As a quick workaround, if ',5' is in any of the column names please try renaming them to remove it.
Thanks Russ,
for the prompt reply!
There is no such thing as ",5" in the dataset. I used the "Reactor 20 Custom" dataset from the DOE library.
Markus
Hi XGBoosters,
Several improvements for XGBoost are available in JMP 17 Pro Early Adopter 6, and require re-installation of the add-in (please download XGBoost.jmpaddin again from this page and re-install). Improvements include the following:
- Monotone and Interaction constraints are implemented as new text-edit boxes under Advanced Options in the dialog. You must type in the appropriate syntax as described in the XGBoost documentation.
- The Pseudo-Huber objective function is available as a new selection in the Objective pull-down menu.
- Shapley values are available under the red triangle for each distinct model fit as the option Save SHAPs. Clicking this option adds new columns to the data table, one for each feature.
- H Measure appears by default for binary targets in the ROC plot output and is also available for display in under the Model Comparison red triangle.
- Survival prediction is available by using the new Censor role in the initial dialog and after choosing the survival:cox objective function in the main dialog.
As always, your feedback is most welcome. If you are a licensed JMP user, you can request access to the JMP Pro 17 Early Adopter Program at jmp.com/earlyadopter .
Hello Russ,
I have experienced a similar issue as Markus described in his previous post when changing from JMP15pro to JMP 16 pro and wondered whether there is any solution to this bug.
Whenever I try to run XG boost on any dataset (including the provided example data) I receive this error:
I have tried to reinstall the addin and JMP several times but could not get it to work. Any ideas? I am on a Windows computer.
Thank you in advance
Greetings
Fred
Hi @fkistner @markus Thanks for reporting. There was a bug in JMP Pro 16.0 that has been fixed in 16.1, although it typically results in a crash and not the error window you mention with ",5". XGBoost models with Reactor 20 Custom and other data sets run cleanly for me in JMP Pro 16.1, 16.2, and 17 Early Adopter using Windows with English language settings. I'm honestly not sure what is triggering the ",5" error you are seeing and have yet to reproduce it. If you can provide more specific details I can investigate further. If you can switch to JMP Pro 17 EA6 that would be best as it has the latest functionality (see note above) and is where I will make further enhancements in preparation for the official 17 rollout later this year.
Hi, Russ
Also I get the Alert message
I did an upgrade to JMP16.2 without any effect
Il will register for the JMP17 early adopter version.
Greetings,
Lu
@Lu @fkistner @markus I think the error may be caused by a locale setting that uses comma instead of a period for the decimal separator. XGBoost has the parameter base_score under Advanced Options with value 0.5, but it can get passed as "0,5" if you have commas turned on. If you would please try under File > Preferences > Windows Specific > Decimal Separator and explicitly set it to a period. I was able to reproduce the error by setting it to comma, so setting it to period may fix things for you.
Hi Russ
You have found the origin of the problem. Thx helping me and many others
Greetings from Belgium,
Lu
Hi Russ,
thx for the help. Even though changing the delimiter lead to another error in JMP 16 changing to 17 ea finally did the job :-)!
Greetings
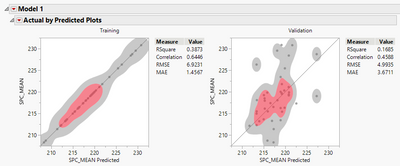
Plot isnt matching RSquare.
@ewittry1 Thanks for reporting. I think this may already be fixed in JMP 17 EA if you want to try it or if you are able to send your data and model to russ.wolfinger@jmp.com I can verify.
Russ,
In other platforms, when I request to generate python code, the program generates ONE python file that works with JMP_SCORE.PY.
When I ran your plugin in JMP 16, I got 3 files instead of 1.
Assuming these three files dont work with jmp_score.py.
Meaning the model inference methodology seems different between XGBOOST and other platforms that generate models.
Since my company builds wrappers around model inference, it would be nice to know if XGBOOST is going to be implemented in JMP_SCORE.PY or what are the correct methods for deployment and inference in JMP 16/17 (same or different). We will have to make accommodations for different platform inference styles.
Also has JMP thought about https://onnx.ai/ , as in something like http://onnx.ai/sklearn-onnx/ , but onnx for JMP?
Hi @ewittry1 , the Python code generated directly from an XGBoost model red triangle is for training the models across all folds from scratch and making out-of-fold predictions. To obtain Python code for scoring/inference like other JMP platforms, first click the model red triangle > Publish Prediction Formula, then from the Formula Depot window, click XGBoost model red triangle > Generate Python code. Please let us know if this doesn't work for you. Thanks for the onnx suggestion; will look into it.
I have been trying to use this add-in and can manage to "Make k-fold columns." But when I actually try to use the XGBoost command, I'm getting the error below. I'm using a Macintosh with M1 chip, and I suspect it is related to that. Any workaround?
'/Users/michaelbailey/Library/Application Support/JMP/Addins/com.jmp.xgboost/lib/15/libxgboost.dylib' (mach-o file, but is an incompatible architecture (have 'x86_64', need 'arm64e'))
Cannot load the XGBoost library. dlopen(/Users/michaelbailey/Library/Application Support/JMP/Addins/com.jmp.xgboost/lib/15/libxgboost.dylib, 0x0002): tried: '/Users/michaelbailey/Library/Application Support/JMP/Addins/com.jmp.xgboost/lib/15/libxgboost.dylib' (mach-o file, but is an incompatible architecture (have 'x86_64', need 'arm64e'))
@profjmb Thanks for notifying us about this, and you are indeed right, we had not yet compiled a libxgboost.dylib for the Apple Mac M1/AMD architecture. I just uploaded an updated version of the add-in that includes this, with kind help from @SamGardner and @chrishumphrey . You're going to need to obtain an early adopter version of JMP Pro 17 for it to work, which you can request at jmp.com/earlyadopter/ .
Hello, I just downloaded JMP 17 EA 8 and XGBoost here. My system crashes (just before completion of autotune) when running autotune on sample data set "powder metallurgy".
thanks
Matteo
Thanks @matteo_patelmo , a fix for this crash should be available in EA9. In the meantime, a workaround is to run at least one model first without autotune.
Thanks, the workaround works.
Autotune is a nice feature, it would be nice to see it implemented in other platforms as well.
Matteo
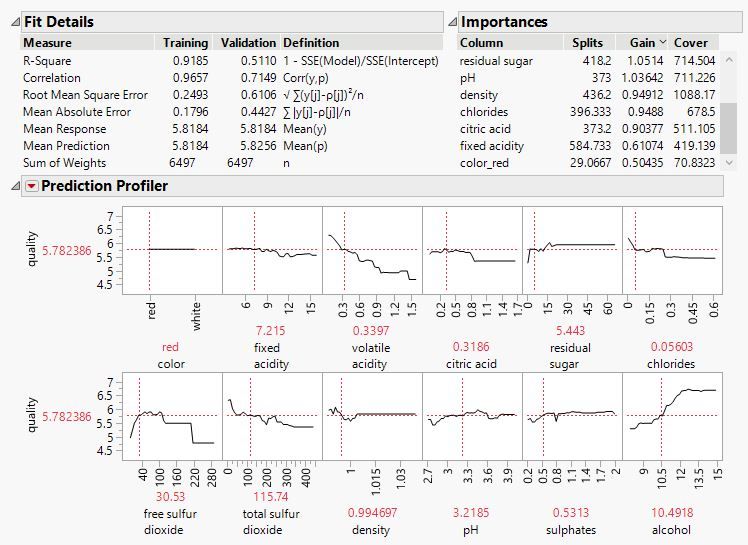
I am trying to use the parameter importance output of the XGBoost to make a selection of predictors for an outcome predicted. However, the paramter importance table of XG boost produces 3 column values: #splits, Gains value and Cover value. Which of these 3 values (#splits, gain or Cover) I use best to prioritize for predictor selction?
regards,
Lu
@Lu , Splits measures the total number of splits on a variable, Gain measures improvement per split, and Cover the amount of data affected by a split. All three provide valuable information about the importance of the predictors. What I usually do:
- Right click > Make into Data Table
- Right click on Feature column > Label/Unlabel
- Select Splits, Gain, Cover in the table, right click > New Formula Column > Log > Log
- Analyze > Multivariate on the logged values
- Study the pairwise scatter plots, mousing over and labeling the points near the upper right corners; these are the most important features
Be aware that simple variable selection on an entire data set can easily lead to overfitting. If you use k-fold cross validation in XGBoost, the importance statistics are averaged over the folds, so this should help mitigate this somewhat. Also, it is safe to drop variables that have zero splits, since they never entered any of the constitutive trees.
The profiler and Shapley values can also be helpful in assessing feature importance.
Thanks Russ,
Instead of using the Multivariate analysis and the graphical way, I calculated the sum via selecting the log transformed columns of (log)#splits, (log)-gain and (log)-cover then New Formula columns > combine > Sum. Then the new Column > Sort > Descending to sort by maximum value to lower value. This way, the column is sorted by a optimal combination of spits, gain and cover value.
Regards,
Lu
Dear Russ,
Is there a way to perform XGBoost together with other ML models in the Model screening platform using the Nested Cross validation option? When I try this option I only get summary statistics for the ML models but not for XGBoost.
Lu
Dear Russ,
I tried to use the model screening function to compare XGBoost with other machine-learning and statistical model. However, for XGBoost, there is no test set data. Why is that? What is the possible way that I can compare the RASE of test data set between XGBoost and other methods?
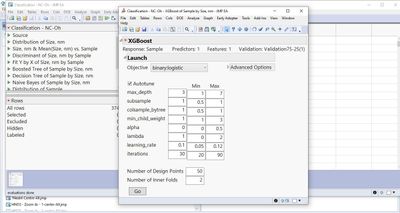
Dear Russ, I have some doubts, which may be due to basic mistake. In JMP Pro 17EA, the XGBoost has a new interface, which made me feel a bit confused. For example, my case is to predict material type from the respective particles size distributions (classification). For example, the Tuning Table currently has got some Inner Folds, which is unknown to me. What to do with the Inner Folds? Which number must be selected in its field?
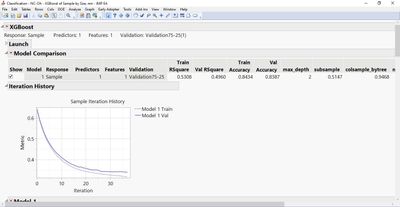
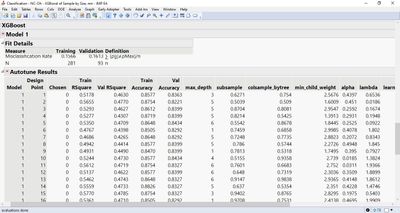
Also, Number of Design Points intuitively speaks for number of models, which I used to encounter after running a code.
But now after the code running, just a single model is exhibited instead of the models list,...
...whereas, the points list is provided to me.
I will highly appreciate your help, if you share the solution with me, what to do if I need to deal with the set of models and how to select the optimal model for me, because I like more the previous version of XGboost Add-in with the corresponding Manual in pdf.
Regards, Michael
Dear Russ,
In JMP17ea XGboost plaform calculates the Shapley value. Do you know whether it is the Kernel or the HyperShap method that is used?
regards,
Lu
@Lu , There is currently no way to get XGBoost output from Model Screening with the Nested Cross Validation option (we may work to remedy this in a future release). A valid approximate comparison would be to run other JMP platforms with Nested CV and then XGBoost by itself with the same number of repeated K-folds (3-5 repeats should be sufficient in most cases). The exact folds will not be the same but the average performance metrics on the test set splits should be comparable.
Shapley values are obtained directly from the XGBoost executable, and I believe they are similar if not exactly the same as the TreeExplainer from https://github.com/slundberg/shap (not Kernel or HyperShap).
@TasapornV XGBoost currently only recognizes training and validation sets, not a third partition for test sets. As mentioned above to @Lu, suggest comparing Nested CV results in Model Screening to the same number of repeated k-folds running separately in XGBoost. I currently do not recommend comparing basic (non-nested) K-fold CV results because several JMP platforms peek at the validation set during training, and so their validation results are overly optimistic. XGBoost does not use the validation data in any way during training and so its K-fold CV results are leak-free; i.e. its validation sets are in fact test sets.
@Nazarkovsky As you have noticed, the new Autotune functionality now selects a single final ensemble model automatically for you over nested k-folds, saving time and protecting against overfitting that could result by manually cherry picking over a large number of runs. It is your choice regarding the number of nested inner folds to use, although it will likely not make too much difference and the default of 2 should work okay in most cases. If you really want separate full model results over a collection of parameter settings, set up the parameter values in a separate JMP table (generated by right clicking a Model Comparison table and modifying parameter values, or by Custom DOE, or by just entering values by hand in a new table with parameter names as columns), and then specify this table in the Tuning Design Table under Advanced Options (type the name of the table in the box).
@russ_wolfinger Dear Russ, thanks for your patience and time!
Another thing: in case of the use of the validation column prepared through "Make Validation Column", what to do with Inner Folds? I mean, if I insert validation column into the "Validation" field while launching the XGBoost platform. Need some time to get used to dealing with the new interface
thanks!
@Nazarkovsky The inner folds are currently automatically generated using a mod function on the row number and you do not have to specify any variables to control them like you do for the outer validation folds. You just need to specify how many inner fold spits to create (default 2 for speed).
Dear Russ,
Not totaly clear to me. Can you make a screen shot of the suggested approximate comparison methods for in case of a classic nested cross validation and in a case of XG boost?
Are paramters in the result tables of the 2 methods comparable (e.g. mean AUC)?.
I am confused about the difference between Fold number and Test set.
Regards,
Lu
Dear @russ_wolfinger,
Great addin ! very useful!
I get an error trying the addin on a Apple chip M2, please see error below:
XGBoost Error: [13:34:50] /Users/runner/work/xgboost/xgboost/python-package/build/temp.macosx-11.0-arm64-3.8/xgboost/src/gbm/../common/common.h:239: XGBoost version not compiled with GPU support.
Stack trace:
[bt] (0) 1 libxgboost.dylib 0x0000000146a83c34 dmlc::LogMessageFatal::~LogMessageFatal() + 124
[bt] (1) 2 libxgboost.dylib 0x0000000146b29b88 xgboost::gbm::GBTree::GetPredictor(xgboost::HostDeviceVector<float> const*, xgboost::DMatrix*) const + 120
[bt] (2) 3 libxgboost.dylib 0x0000000146b2b360 xgboost::gbm::GBTree::PredictBatch(xgboost::DMatrix*, xgboost::PredictionCacheEntry*, bool, unsigned int, unsigned int) + 228
[bt] (3) 4 libxgboost.dylib 0x0000000146b50454 xgboost::LearnerImpl::PredictRaw(xgboost::DMatrix*, xgboost::PredictionCacheEntry*, bool, unsigned int, unsigned int) const + 108
[bt] (4) 5 libxgboost.dylib 0x0000000146b40e64 xgboost::LearnerImpl::UpdateOneIter(int, std::__1::shared_ptr<xgboost::DMatrix>) + 420
[bt] (5) 6 libxgboost.dylib 0x0000000146a879a4 XGBoosterUpdateOneIter + 140
[bt] (6) 7 JMP 0x00000001052f8ef8 getLegendModel(DisplayBox const*, int) + 214016
[bt] (7) 8 JMP 0x00000001052d9de8 getLegendModel(DisplayBox const*, int) + 86768
[bt] (8) 9 JMP 0x00000001052e2a9c getLegendModel(DisplayBox const*, int) + 122788
@Vins Thank you for the error message on Apple M2; we will look into it. Suggestions in the meantime:
1. If you have gpu_predictor selected under Advanced Options, it might work to change it back to the default value of cpu_predictor (still should run very fast).
2. It might work to find M2-compiled versions of libxgboost.dylib and libomp.dylib and drop them into the add-in folder to replace the current versions (which were compiled only for M1 and previous chips). You can find these two files if you install XGBoost in Python then search into the Python environment install directory. To find the add-in file Home folder, click View > Add-Ins, click on XGBoost, click on the home folder link, then look in ../lib/17/. Make sure to save backups of the current files before doing this.
Dear Russ,
i have some Problems with the Autotune function. It only creates one more Model at a time and not multiple.
It still worked with version 16.
Now I have updated JMP Pro to version 17/17.1. I reinstalled the add-in. Now when I use the autotune function here, only one more model is added for each time I click Go. Fit X/10 Model is also displayed in the loading window.
It also doesn't work on the Powder Metallurgy sample file.
Do you have any advice? Are there two different versions of the add-in? (V16 and V17?)
Another question: is there a possibility to use the autotune function for the advanced options or objectives in XGBoost?
Best regards
Stefan
Hi @StefanS, This new way of doing Autotune is intentional, as the previous way of outputting the separate model fits tends to promote cherry-picking and overfitting. At the bottom of the new single model fit you should see a table of results that can help you settle in on good hyperparameters. If you want the previous behavior, there is a Tuning Table option under Advanced Options, for which you must specify a separate JMP table of parameter values, one row per setting and columns corresponding to the parameters you want to tune. These can include other advanced option parameters like you asked about. You can create this table with DOE (Custom or Easy) or by hand.
Hi @russ_wolfinger ,
A while back you had a post on use of xgboost for time series analysis. Is there a link to the post? I was keen to see how you used xgboost for time series and how it performed vs other time series methods
@Vins Thanks for raising this and reminding me to cross-post my note on time series forecasting with XGBoost. For some reason this appears to be difficult to find with the search bar. Here is the link.
Being classically trained in both time- and frequency-domain time series methods like ARIMA and periodograms, I remain amazed at how well boosted trees perform with forecasting. Deep neural nets are likely overtaking them now in many cases (please register for JMP 18 Early Adopter to try the forthcoming Torch Deep Learning add-in), or at least provide a great complement to boosted tree models for ensembling.
Hi @russ_wolfinger,
The link is not active, I get a "access denied" error message.
I am keen to try the Deep Learning add-in! I have registered for the early adopter program!
Hi XGBoosters,
If you are interested in seeing a live demo of using XGBoost for time series forecasting in both Python and JMP Pro, please check out a free online webinar by @kemal and me on 20 June at 2 PM EST in an event hosted by Wiley. We have over 400 registered so far and would love to see you there. Register at https://events.bizzabo.com/487993
Just posted! Webinar series shows attendees how to conquer the 'do more with less' paradox. Includes materials from the aforementioned webinar.
- © 2024 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- About JMP
- JMP Software
- JMP User Community
- Contact