- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
JMP Add-Ins
Download and share JMP add-ins- JMP User Community
- :
- File Exchange
- :
- JMP Add-Ins
- :
- The JMP to R Add-In Builder
Update 27Jun2018: JMPtoRAddinBuilder.jmpaddin now contains unencrypted JSL source files.
Version 2 Update
I am happy to announce that the JMP to R Add-In Builder now supports the output of all output components (table, plot, etc.) as a single report! To enable this feature, install the attached "JMPtoRAddinBuilderWithReport". When defining your output components, check the "Return all results as report" box.
To see an example of the updated output format, scroll to the corresponding section in the post below. The SVM function with the report output enabled is also attached.
What is the JMP to R Add-In Builder?
While JMP is a very powerful statistical software, JMP users may want to extend JMP beyond its included capabilities by accessing some R functionalities.
This summer, in light of the need to make R functions more accessible to JMP users without leaving the JMP platform, I created this guided configuration JMP user interface that allows users to define and use custom JMP add-ins that execute R functions. This add-in allows end users to analyze their data in JMP using R functions without writing JSL or R code.
This program utilizes the JMP to R connection, and can also be adapted to create custom add-ins for functions in other languages, such as Python and JSL itself.
See a video demo of the JMP to R Add-In Builder here or below:
Requirements
- R must be installed on your computer. Install R from https://www.r-project.org/
- The add-in only runs on Windows OS.
- If applicable, install the appropriate packages in R that you will use in your R code.
Usage Notes
- If you use column selects or column lists in your add-in, be sure to open your data table prior to running the configuration UI.
- Each function created will only have one column list maximum.
- All the column selects will be grouped together in one column; enter the same column for all column selects.
- Each of the button names for column selects must be unique.
- Column selects and column lists will be placed at the top of their respective column.
- In your R code, reference your data table as “dt”.
- If you are appending a column to a table and are having the user select the column type, in your R code, refer to the string that contains the user’s choice as “colType”.
- The values for check boxes are returned as ‘0’ (unchecked) or ‘1’ (checked).
- Paste R code in as you normally would in the R console except for the following difference.
When referencing the name of a variable as a string as a parameter of a function, use the eval(parse()) function in R. For example:
This evaluates the expression and uses the string stored in colnames(factor)[1] in the function operation.eval(parse(text=paste("x <- subset(dt, select=-", colnames(factor)[1], ")")))
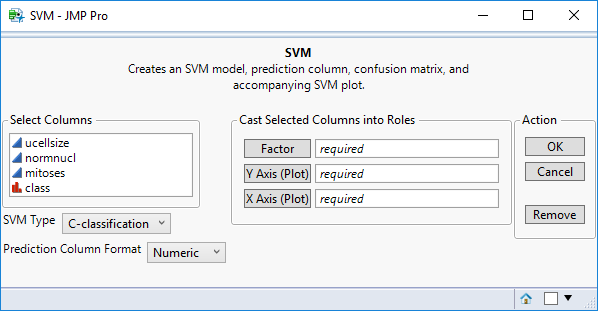
How It Works: Creating a Support Vector Machine (SVM) Function as an Example
STEP 0
Install and open the JMP to R Add-In Builder.
NOTE: If you use column selects or column lists in your add-in, be sure to open your data table prior to running the configuration UI AND prior to running the add-in you create.
STEP 1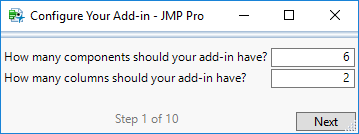
*The five types of components supported are: Check Box, Column List, Column Select, Dropdown Menu, and Numeric Value. Each column select comes with its own assignment button, so count each column select and corresponding button as one component.
#The number of columns refer to display columns in which components appear. For example, there are two display columns in the UI for this example (see below).
NOTE: The “Action” column is grouped with the column select column, meaning those two objects grouped together counts as one column.
STEP 2
Next, for each of your components, identify the type.
NOTE: Enter the components in the order you’d like them to appear on your add-in, column-wise. This means that all components in each column must be in sequential order in the dropdowns you specified, but you do not have to enter values strictly column by column.
For example, you can enter the types for component 1, column 1; then component 1, column 2; then component 2, column 1; then component 2, column 2 rather than column by column if you so desire.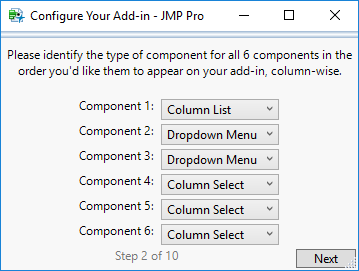
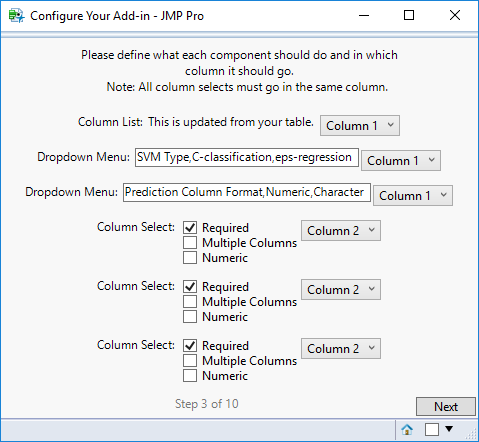
STEP 3
For each component, select the column in which it should appear.
On this screen, also define the parameters for each component as follows:
- Check Box: Enter the name (label) for the component.
- Column List: Other than selecting the column, no action needed.
- Column Select: Check all the properties that apply to your selection box.
- Dropdown Menu: Enter the label for the dropdown menu and the choices for the menu. DO NOT include a space after commas (i.e. list things as a,b,c,d), but the names and choices themselves can contain spaces. The first name you write in the list will be taken as the name for the dropdown menu, and the rest will be listed as choices in the order in which you type them.
- Numeric Value: Enter the name (label) for the component.
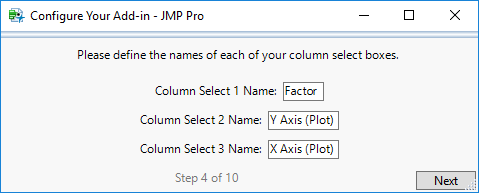
STEP 4
If your add-in uses column selects, enter the names for each of the buttons for your corresponding column selects in the order they should appear. Otherwise, click ‘Next’.
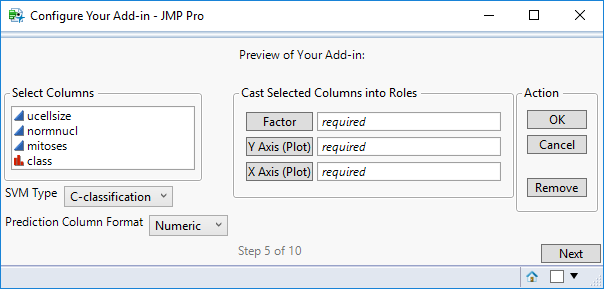
STEP 5
This screen displays a preview of the add-in UI you’ve created.
STEP 6
On this page, name your add-in and give it a description (which will be displayed under the add-in name).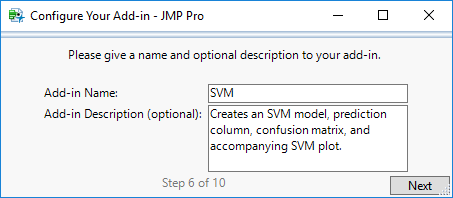
STEP 7
On this screen, define the input and output variable names that will be used in your R code. These should only be one word (no spaces).
For the output variables, DO NOT include a space after commas (i.e. list variables as a,b,c).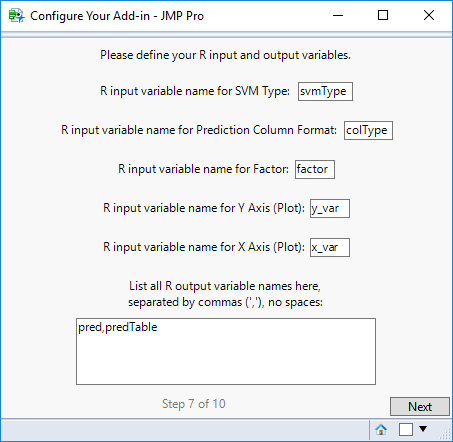
STEP 8
On this screen, enter the R code that your function will run. The R code should be in R syntax (i.e. could be run directly in an R console).
The code should reference the variable names you declared on the previous screen.
Also enter the number of output displays for the function. The four types of output displays currently supported are:
- Append Column in Existing Table
- Most Recent R Plot
- Open Table
- Results Window with Text or Numbers
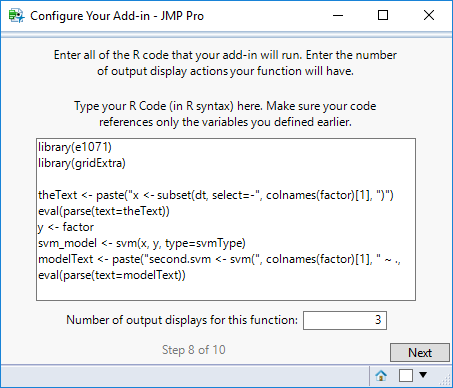
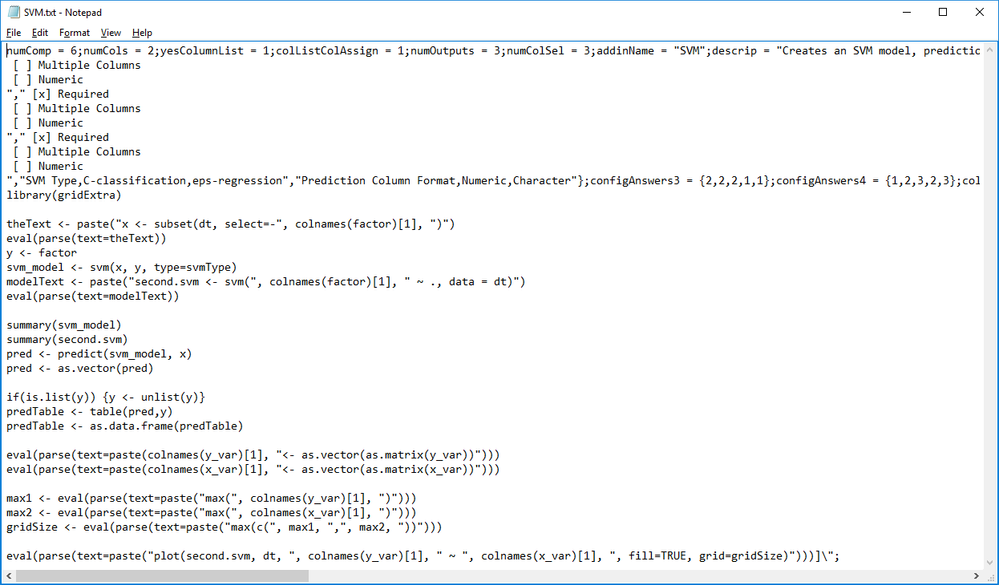
R Code for this example:
library(e1071)
library(gridExtra)
theText <- paste("x <- subset(dt, select=-", colnames(factor)[1], ")")
eval(parse(text=theText))
y <- factor
svm_model <- svm(x, y, type=svmType)
modelText <- paste("second.svm <- svm(", colnames(factor)[1], " ~ ., data = dt)")
eval(parse(text=modelText))
summary(svm_model)
summary(second.svm)
pred <- predict(svm_model, x)
pred <- as.vector(pred)
if(is.list(y)) {y <- unlist(y)}
predTable <- table(pred,y)
predTable <- as.data.frame(predTable)
eval(parse(text=paste(colnames(y_var)[1], "<- as.vector(as.matrix(y_var))")))
eval(parse(text=paste(colnames(x_var)[1], "<- as.vector(as.matrix(x_var))")))
max1 <- eval(parse(text=paste("max(", colnames(y_var)[1], ")")))
max2 <- eval(parse(text=paste("max(", colnames(x_var)[1], ")")))
gridSize <- eval(parse(text=paste("max(c(", max1, ",", max2, "))")))
eval(parse(text=paste("plot(second.svm, dt, ", colnames(y_var)[1], " ~ ", colnames(x_var)[1], ", fill=TRUE, grid=gridSize)")))
STEP 9
For each of the output displays you have selected, specify the type.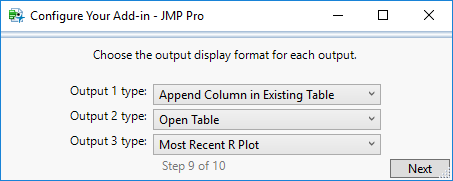
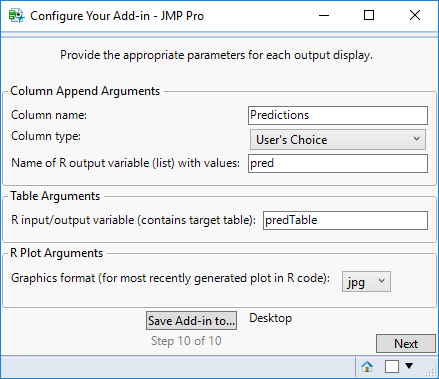
STEP 10
Provide the parameters for each output display you selected as follows:
- For Column Append, enter the name of the column that will be appended to the currently open table, the type of data the column contains, and the output variable specified earlier that will contain the list of values that will populate the column.
- For Table, enter the output variable specified earlier that will contain the table.
- For R Plot, enter the format of the graphics you want to retrieve from R (png or jpg).
- For Window, enter the results text as it should appear in the results window. If you would like the values of input/output variables to be displayed, surround the variable names with the caret (^) symbol. Also enter the name of the window you’d like to be displayed.
To install the add-in, double-click on the add-in and JMP should ask you whether to install it. Click “Install”. It should now appear in your Add-Ins menu in JMP.
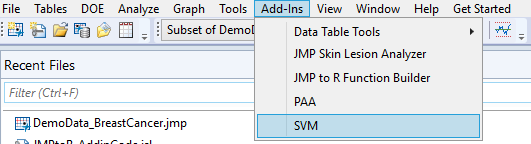
To run the add-in, open the data table you’d like to perform the operation on (if applicable) so that it is the current table. Click on your add-in as shown above and the add-in dialog should open.
For this example, the three outputs are a prediction column, confusion matrix, and SVM plot. The results for the inputs shown above are as follows, with the first view being the newly added report output format in version 2: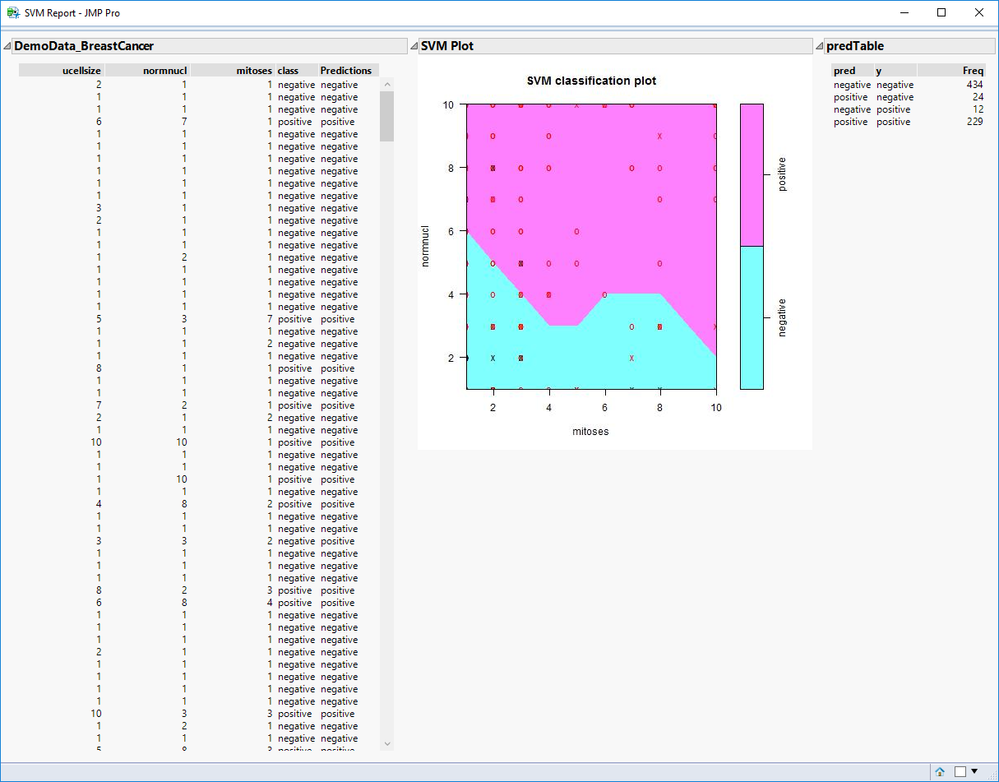
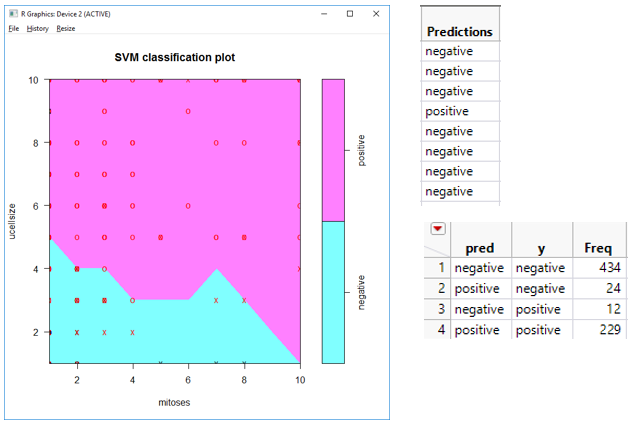
Editing Add-in
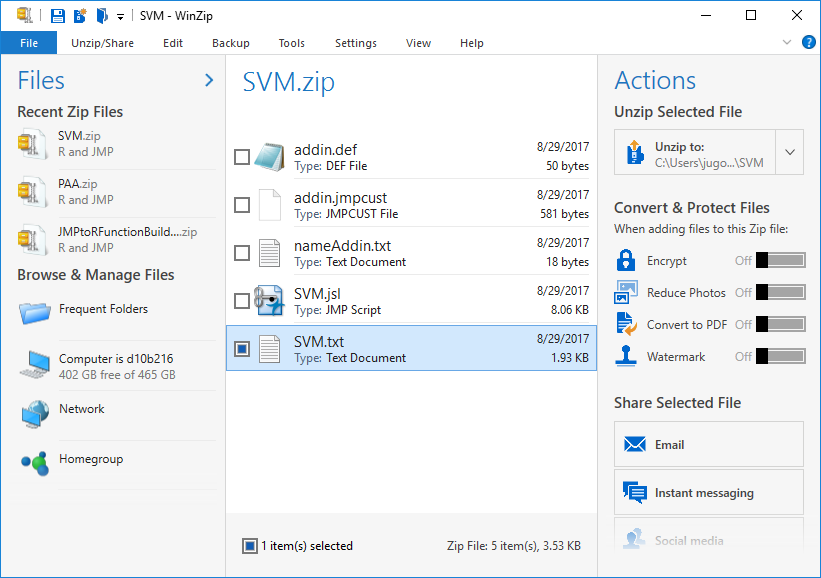
To edit your add-in, open the directory in which you saved your .jmpaddin file. Change the extension to .zip:
Open up the .txt file that has your add-in’s name:
This is your config file. If you wish to make any changes to your add-in, such as R code or variable names, you can directly edit your inputs in this document, save the .txt file and the .zip file, change the extension back to .jmpaddin, and reinstall your add-in. The changes will be reflected if you changed the values correctly.
Please see the attached documentation for a second example (Piecewise Aggregate Approximation of Time Series, R code adapted from here) and other notes. Attached are also the two examples (SVM, PAA) I created with this add-in.
What R add-ins have you created with this add-in? What improvements would you like to see? Let me know in the comments below!
Thanks for the "JMP to R Add-In Builder" !
Are you planning to create a Mac version? After all, the "M" in JMP stans for a lot of users, doesn't it?
Cheers,
@Alfredo -- this was a summer project completed by one of our interns and we ran out of summer before completing everything that we wanted to (including macOS support). Hopefully we can pick this back up next summer!
- Mark as Read
- Mark as New
- Bookmark
- Get Direct Link
- Report Inappropriate Content
This is great, particularly for those of us like me who are non-coders. Thanks fo your contribution.
I'd like to second the Mac version request.
This looks like a life-changing feature, but I'm confused about how to use it. My attempt doesn't work, and I don't know why. Most likely, I'm confused about how to pass information from JSL to R or back. Can you please explain the
eval(parse(text=...))statements? Perhaps it would help me if you colored the text in your example by which functions were JSL and which were R?
If you refer to Usage Note #9, you'll see that eval(parse... is an R function. Also near there you'll see that this only works on Windows OS, so if you're running under OSX, you might get a little frustrated - add your voice for OSX support.
There are other articles demoing how to run R under JMP. If you're needs are simple, you might consult these - passing R code and parameters under JMP is pretty straightforward.
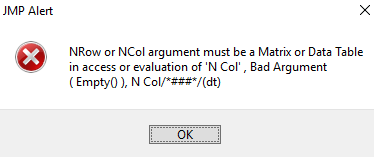
The new version (add-in without encryption) has this error when I get to step 4:
Regarding the NRow or NCol error above, if anyone else finds this error, I think I found the cause. A bug in the JSL code generated by the addin builder near line 140. I think the line:
createdAddin[4][configAnswers3[p]][1][configAnswers4[p]][2] << get()should read
createdAddin[4][configAnswers3[p]][1][ notColSelectCt + yesColumnList][2] << get()This is brilliant! JMP, please assign a new summer intern to expand this to Mac.
I see this "can also be adapted to create custom add-ins for functions in other languages, such as Python". How would we create a plugin that uses python?
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us