- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
JMP Add-Ins
Download and share JMP add-ins- JMP User Community
- :
- File Exchange
- :
- JMP Add-Ins
- :
- Spectral Tools
Description
Spectral Tools is a collaborative effort of @JeremyAshJMP and @Bill_Worley . It provides some convenience functions for analyzing spectral data, or related data (chromatograms, etc.). Most of the functionality can already be performed in JMP, but the intent was to streamline the process. Hopefully this will help lower the barrier to entry for new users, and make routine tasks faster for experienced users. See part 1 and part 2 of our blog describing the spectral analysis workflow.
Usage
Example Data: An example spectral data set from Martens et al. Anal Chem. 2003 Feb1;75(3):394-404.
Import Spectra: When importing spectra, Spectral Tools assumes that each spectra is in a separate file in a single directory.
Import Spectra > Import Delimited Text Files: Spectra can be in delimited text files (.csv or .tsv, for example). Spectral Tools assumes the X variable is on the first column, and the Y variable is on the second.
Import Spectra > Import JCAMP-DX Files: The simplest variation of JCAMP-DX files is also supported. This means JCAMP-DX files without compression and with a XYDATA=(X++(Y..Y)) variable list. I have not tested this out on different variations of the file format. I plan to add to this post some data that I did test with, once I get permission from the owners.
Preprocessing > Standard Normal Variate: Makes a new column with the standard normal variate applied to the spectra. Assumes your data is in the stacked format.
Preprocessing > Savtizky-Golay Smoothing: Launches Graph Builder with a Savitzky-Golay smoother applied to each spectra. Select your smoothing parameter (local width) and then use the button at the top left of the report to make a new column for the smoother. Assumes your data is in the stacked format. Only available in JMP 16 and higher.
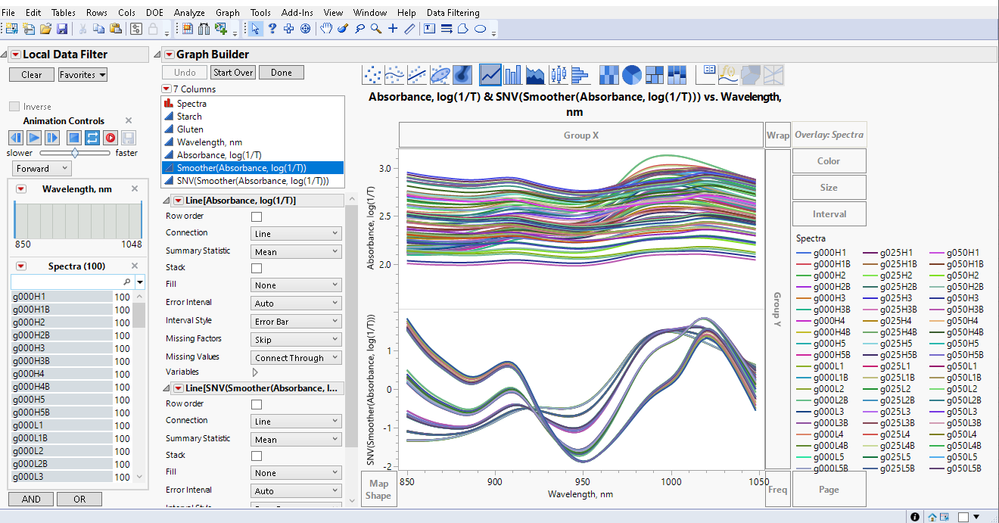
Launch Graph Builder: Launches a spectra line plot for one spectra column or two spectra columns for comparison (e.g., before and after pre-processing). Some of the features you might want to use are set up for you. Assumes your data is in the stacked format.
See this blog post for a Savtizky-Golay derivative filter addin and demonstrated usage.
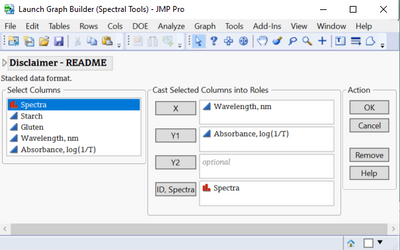
Disclaimer
There is a disclaimer that appears in each launch dialog. Basically it says that this is only a prototype. This has not been rigorously tested, and does not do much in terms of error handling. We do not intend to provide regular maintenance or support, and this is only a first step towards making analyzing spectra more convenient in JMP.
That being said, it would be great if you could try it out and give us feedback in the comments, or by message to @Bill_Worley. We probably won’t have time to implement new features in the add-in, but it would be helpful for development to hear about features you would like to see in the future. The JMP Wish List is also a great place to make requests.
Thanks you two! I was always missing better tools for spectral analysis.
Unfortunately, trying to use the launch graph builder with the sample data crashes my JMP 15.2
Thanks @Feli. Well that wasn't very graceful, was it? Looks like there is JSL in the addin that is 16 dependent. I didn't realized that 16 hadn't shipped yet. It should be available late March, I believe. I am working to make this backward compatible with 15.2. I hope to have a fix up soon.
@Feli, the attached update should work with 15.2. Note that the Savitsky-Golay smoothing is only available in 16. I tested the addin for backward compatibility through JMP 14. Thanks!
would be useful to build also some spectral data deconvolution capability
Will you be able to give us option when importing files in the folder to choose which line will start the data for X & Y variables?
Our spectral data for FTIR raw data format has 37 rows and 2 columns of file description. So Column#1 will have character labels, Column#2 will have mixed values of characters and numbers. After then, all rows as of row#38 on column#1&2 will have numbers in scientific figures format.
So in other words, can we truncate all data above specific row number and stack data including file names when importing?
Below is an example of this.
| RES | 4 |
| APERTURE | OPEN |
| BEAM | INT |
| DET | INT1 |
| FILTER | 1.2 |
| SENSITIVITY | 1 |
| SYM | SYM |
| UDR | 2 |
| SCANS | 64 |
| XAxisLabel | Wavenumber |
| YAxisLabel | Absorbance |
| Title | Absorbance |
| Point Separation | 1.93E+00 |
| Starting Point Number | 207 |
| Peak Location | 3555 |
| XLabel | 20402 |
| YLabel | 20502 |
| FileType | 20602 |
| DisplayDirection | 20300 |
| PeakDirection | 20312 |
| X Position (mm) | 0 |
| Y Position (mm) | 0 |
| Radius (mm) | 0 |
| Theta (deg) | 90 |
| Site Number | 1 |
| Time | 06302021_155524 |
| Recipe | asdadsadsd |
| Background | |
| Scan Speed (kHz) | 5 |
| Baseline Method | Local/Piecewise/Linear |
| Smoothing Used | FALSE |
| Smoothing Method | Moving Average |
| Smoothing Window Length | 3 |
| Smoothing Polynomial Order | 2 |
| Atmospheric Compensation Used | TRUE |
| CO2 Reference Spectrum | ReferenceSpectrumCO2_20210617T084605.csv |
| H2O Reference Spectrum | ReferenceSpectrumH2O_20210617T084605.csv |
| Wavenumber | Absorbance |
| 3.99263973E+02 | 4.29913431E-02 |
| 4.01192784E+02 | 4.45658502E-02 |
| 4.03121596E+02 | 4.56853577E-02 |
| 4.05050407E+02 | 4.88489802E-02 |
| 4.06979219E+02 | 5.18775801E-02 |
| 4.08908030E+02 | 5.25619803E-02 |
| 4.10836841E+02 | 5.26030929E-02 |
| 4.12765653E+02 | 5.19562901E-02 |
| 4.14694464E+02 | 5.12063073E-02 |
| 4.16623276E+02 | 5.23197525E-02 |
@joelahn Sorry about the delayed response. I am assuming the data is saved in a delimited text file, such as a csv? For delimited text files, spectral tools just provides a streamlined interface to the multiple file importer, which you should be able to get to work for your task. Would you try the following:
1. Go to File >> Import Multiple Files.
2. Go to the directory of files and select the files you would like to import.
3. Check add file name column, if you want a file name column.
4. Go to settings. In the had header section. Set the first data line to 38. Your header line is on line 37 and is one line.
5. Click import.
That procedure worked for me in a test data set I just mocked up. It should be easy to add this option into the addin, so if this works for you I will go ahead and do that. Thanks for the request and the clear example of what you need!
When working with the SpectralTools recently (so much fun now, thank you!), I was missing two features:
- easy subtraction of a reference spectrum (e.g. water)
- I like to jump off directly from the spectra to a PLS. Unfortunately, PLS only works on wide table format and not the stacked one, unlike FDE. A button to create the correct unstacked format would be nice
The possibility to add reference concentrations to the spectra (y variable in the PLS) would also be nice
@Feli Thanks for this feedback. Any ideas you have on ways we can improve the workflow would be helpful!
Subtracting off a reference spectrum would be easy to add as a preprocessing option in the addin, so I will try to get around to that soon.
I will pass your other suggestion about PLS onto development. We have been considering doing something like that.
I am not sure I understand your last request, though. Could you clarify what you mean by add reference concentrations to the spectra?
@Feli @Vincenzo Could I get some clarification on what you are looking for as far as deconvolution? This often refers to 1) detecting peaks and computing peak parameters such as peak areas, or 2) fitting a full fledged mixture model to spectra and estimating the underlying mixture components. 2 would be more challenging and is often a tedious process in other software because it requires a lot of tweaking of parameters in the mixture fitter. But it would be good to know if there is a lot of user interest in this....
Hi Jeremy,
In the past, I have analyzed EPR spectra doing something in between 1) and 2). I was fitting the curves using a fixed number (defined a priori) of Gaussian or Lorentzian functions. JMP was behaving well with stable fit with some educated initial guessing.
I was also fitting SAXS and UV VIS curves, but for those the mixtures model were not really useful, I was decomposing curves using PCA as first step, the I was creating better decomposition to fit kinetics odel using multivariate curve resolution, however this later step was not possible in JMP.
In general chemometrics is a very interesting fields with many methods available and with interesting application. I was specifically interested in analysing a series of spectra generated varying temperature and time and I was attempting to create pseudokinetics models, understanding how many independent phenomena were undergoing at the same time.
@Vincenzo Mixture modeling of EPR spectra is a perfect example of an application for 2 we were thinking about. Really helpful to hear that you were able to get JMP to fit mixtures of peaks with some educated guessing in nonlinear.
It sounds like you think both 1 and 2 would be useful, as 1 will often give you educated guesses you need to get a good solution for 2. I know people that have modeled similar type of spectra (XRD, for example) and had a lot of difficulty getting the educated guesses necessary to get a good fit. You are often having to try out different numbers of components, initializations, constraints. So my read on 2 is that making something that would work well in general might take a rich UI with a lot of opportunity for user intervention. Let us know if you feel differently though.
We are definitely interested in making both 1 and 2 more user friendly, and it is good to know about your use cases.
Also interesting to hear about your uses case for MCR because this has been requested by other users as well, so I will add this to our record of requests!
Really appreciate it keep the suggestions coming!
thank you very much, looking forward to seeing this implemented
@JeremyAshJMP with adding reference concentrations I mean that you would give a column (or several) to the Addin that contains the concentration of a chemical etc that you are trying to analyze via their spectra. The ID,spectra column is a good start, but I sometimes have more than one reference value and maybe the spectra have their own sample name. Or if I use the concentration as ID column and have more than one spectra with same concentration but different measurement...
Concerning deconvolution, I also think that both 1 and 2 would be a great benefit. MCR is great when you have time resolved data or otherwise ordered, 1 might work better when trying to resolve a single spectrum or (timely) disjointed ones. Peak fitting is a big classic when dealing with spectra.
and what I forgot as preprocessing option: the classic mean centering and scaling would sometimes also be nice. Normalization (to area or peak) can also help.
I'm having trouble importing Raman spectra as text files and doing spectral analysis in JMP. I have been able to use the Functional data explorer to plot and perform baseline correction. However, all of this has been done without being able to add the wavenumber to the plots. I'm thinking of doing some of this in matlab but I feel like I should be able to do it JMP as well. does anyone know how to do this?
thank you.
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us