- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- tablet dissolution profile
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
tablet dissolution profile
Hi,
I have a set of tablets in which one component determines how slowly the active is released (measured at 1, 2, 3, 4, 5 and 6hours).
If I have the results for tablets containing 10, 15, 20 and 25% of this component how can I create a model that will tell me how much of that component is needed to obtain a target profile (target release values at target timepoints)?
Thanks
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: tablet dissolution profile
Also, you might try saving the model and creating a new response to optimize. A good choice would be the root mean square error (RMSE). A column formula could compare the predicted response with the target response using something like this
Sqrt(
Sum(
(P(t1,X) - T(t1))^2 +
(P(t2,X) - T(t2))^2 +
(P(t3,X) - T(t3))^2 +
(P(t4,X) - T(t4))^2 +
(P(t5,X) - T(t5))^2
)
)where the P(t,X) is the fitted model of the predicted response at time t and excipient level X. T(t) is the target level at time t. You could use the Graph > Profiler with this column to optimize X (minimize response with desirability function).
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: tablet dissolution profile
One JMP platform you may want to take a look at is the Fit Curve platform for the type of problem you are trying to address. Here is the link to the JMP online documentation for the Fit Curve platform.
https://www.jmp.com/support/help/13-2/Fit_Curve.shtml
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: tablet dissolution profile
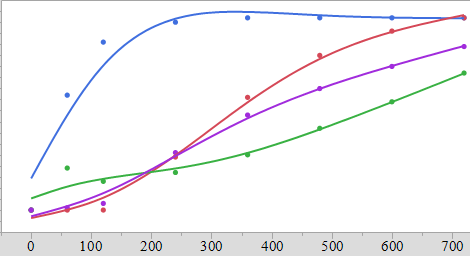
I have no problems with fitting the curves. My problem is that each concentration/fit has a different slope/equation and I'd like to have a model that would tell me what concentration of the excipient is needed to obtain a given profile – that is to say I have a target fit of curve and I'd like to know how I can achieve that based on the results of the other fits.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: tablet dissolution profile
I guess I'm a bit confused then. For your example, are you trying to predict any one curve as a function of the slope/intercept of the other 3 curves?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: tablet dissolution profile
You might also try using Analyze > Reliability > Degradation. Use the excipient level as X. You can try transforms on Y and X with a linear path or use a nonlinear path.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: tablet dissolution profile
I probably do not fully understand how the degradation model works but...
When I have a target profile it means that I have target values for the various timepoints (e.g. timepoint 100: 10%, 200: 15%, 400: 45%, etc.) so not just one point. And I would need a model that would tell me what concnetration of X would give me a closest match.
My best shot so far was to create a single factor DOE where I set the timepoints as the responses and simply overwrote the parameters in the provided table with the parameters and results of my experiments and used the profilers.
But there must be a better way to do that, I guess.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: tablet dissolution profile
Please don't be put off by the terminology. The 'degradation path' in the Degradation platform is just a model of the change in the response (dissolution) over time under the influence of a factor (excipient level). You can then profile the response under different conditions to find the target profile. You can use inverse prediction to find the condition that achieves a desired response level, too.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: tablet dissolution profile
Also, you might try saving the model and creating a new response to optimize. A good choice would be the root mean square error (RMSE). A column formula could compare the predicted response with the target response using something like this
Sqrt(
Sum(
(P(t1,X) - T(t1))^2 +
(P(t2,X) - T(t2))^2 +
(P(t3,X) - T(t3))^2 +
(P(t4,X) - T(t4))^2 +
(P(t5,X) - T(t5))^2
)
)where the P(t,X) is the fitted model of the predicted response at time t and excipient level X. T(t) is the target level at time t. You could use the Graph > Profiler with this column to optimize X (minimize response with desirability function).
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us

