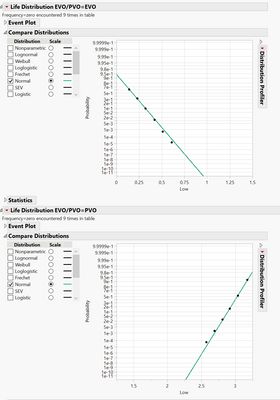
I have an interval censored data table, when I do Life distribution --> compare groups, it gave me Plot1; what I really want is Plot2, in order to do that, I have to do life distribution without compare groups, generate plot3 with separate EVO/PVO plots, change the EVO to show survival curve, manually copy frame contents of PVO plot and paste frame contents into the EVO plot to get plot2. it is very manual, is there a good way to use scripts to generate Plot2?
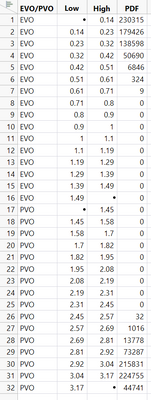
life distribution --> Compare groups
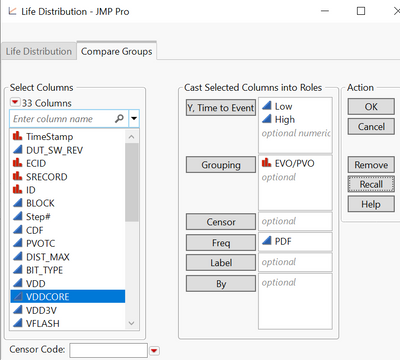
Plot1
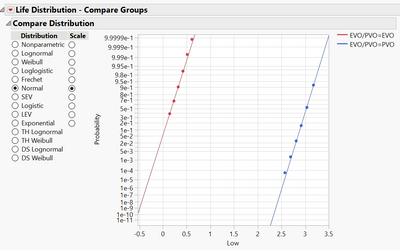
Plot2, left line is EVO with show survival on, right plot is PVO no show survival on;
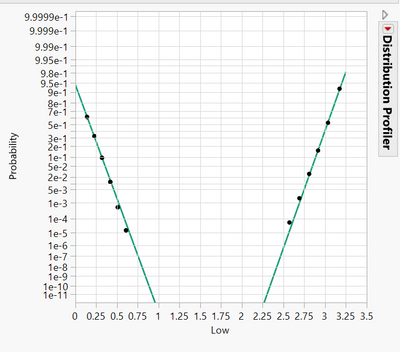
life distribution --> by EVO/PVO
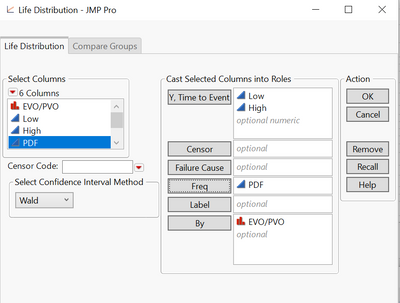
Plot3, EVO has show survival on, PVO has show survival off