- New to JMP? Let the Data Analysis Director guide you through selecting an analysis task, an analysis goal, and a data type. Available now in the JMP Marketplace!
- See how to install JMP Marketplace extensions to customize and enhance JMP.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- How to show fit line graphs in a single report, without the tables and line lege...
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
How to show fit line graphs in a single report, without the tables and line legends
I need to have a quick glance of variables Y1, Y2, Y3, Y4, ..., YN vs a single variable X.
What I do is as follows:
- Fit Y by X simultaneously for all Ys.
- Group by a grouping column G.
- Fit Line function.
- Set the frame size to {150, 125}.
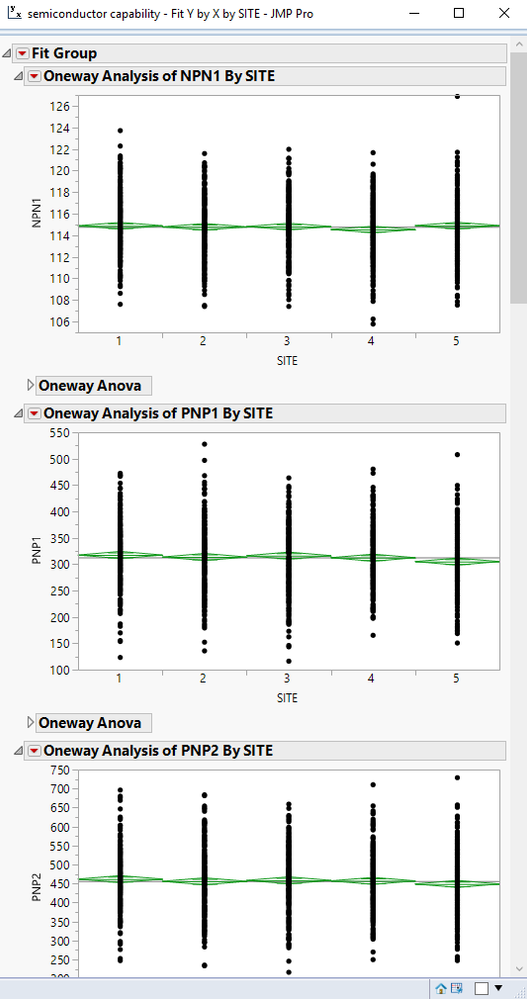
Unfortunately, since I have many cells and Y variables, only two graphs show at a time in a JMP report and I have to scroll several times. What I want is a quick glance for all parameters and probably just scroll once or twice only. In short, I only need the graph itself in the JMP report, no need for the interactive line legend underneath or the ANOVA tables. Is there a way to remove them from the report?
Thanks.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How to show fit line graphs in a single report, without the tables and line legends
There are several things you can do to get the results you want.
1. Close(Collapse) the Anova Results. Using the gray triangle to the left of any of the various sections within the output, you can collapse the details of the section, down to just the Title box itself. If you use the JMP Standard feature of holding down the CNTL key as you click to collapse the Anova segment for one of the parameters, all of the same Anova sections for all of the parameters will be collapsed.
2. Click on the edge of a chart and reduce it in size. Once again, if you hold down the CNTL key as you click on the edge of a graph and change the frame size of the graph, all of the graphs will be changed at the same time.
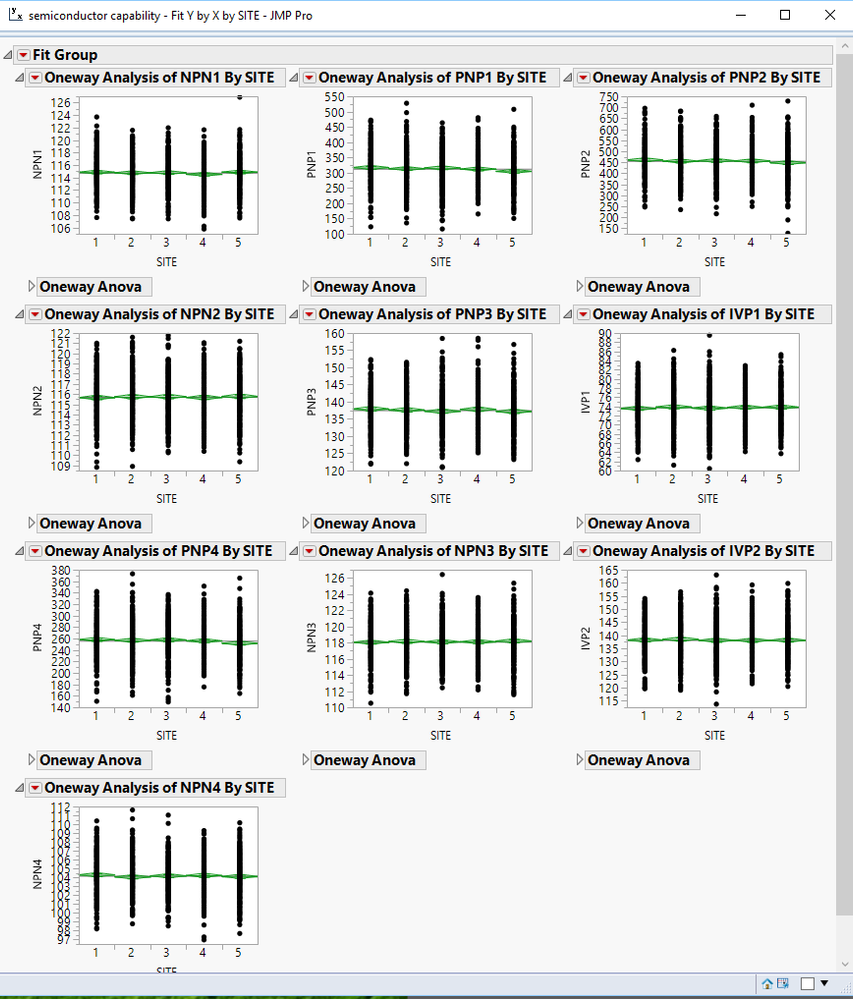
3. If you click on the red triangle for the overall Title for the display, you can select "Arange in Rows". This will allow you to create a matrix of the output. If you have closed all sections except the graphs for each of the parameters, then you will get a matrix of the graphs.
4. A script can be written to modify the displayed output in just about any form you want. Individual components of the output can be deleted, moved, or annotated. The various objects can be placed into new windows and positioned in whatever form you desire.
Names Default To Here( 1 );
dt = Open( "$SAMPLE_DATA/Semiconductor Capability.jmp" );
Fit Group(
Oneway(
Y( :NPN1 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :PNP1 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :PNP2 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :NPN2 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :PNP3 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :IVP1 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :PNP4 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :NPN3 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :IVP2 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
Oneway(
Y( :NPN4 ),
X( :SITE ),
Means( 1 ),
Mean Diamonds( 1 ),
SendToReport(
Dispatch( {}, "Oneway Plot", FrameBox, {Frame Size( 182, 140 )} ),
Dispatch( {}, "Oneway Anova", OutlineBox, {Close( 1 )} ),
Dispatch( {"Oneway Anova"}, "Analysis of Variance", OutlineBox, {Close( 1 )} )
)
),
<<{Arrange in Rows( 3 )}
);
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How to show fit line graphs in a single report, without the tables and line legends
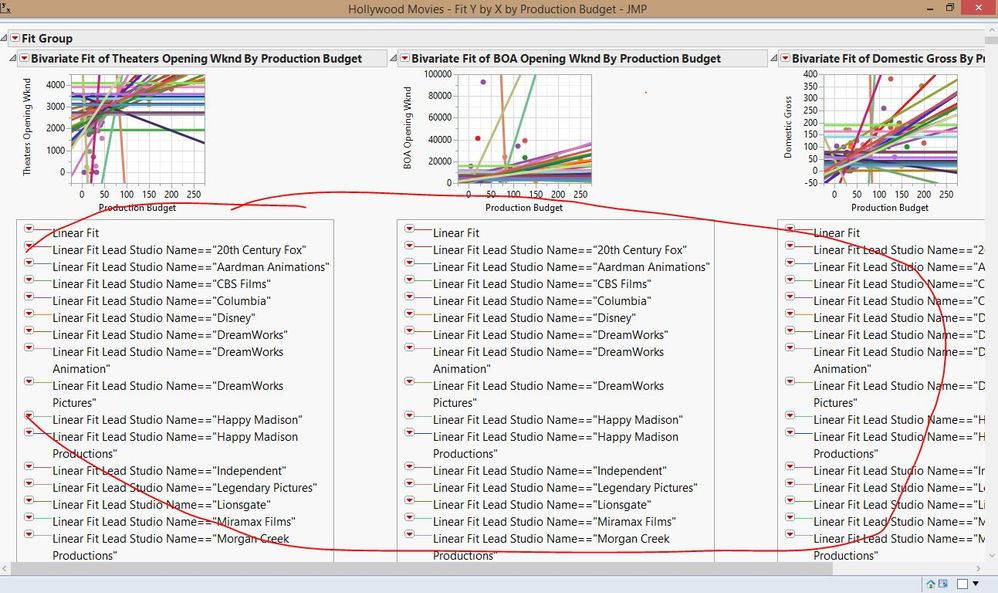
Thank you very much for your comprehensive reply. I can collapse the tables for ANOVA. However, my real problem are the line legends underneath. For example this data from Hollywood movies.jmp (see picture attached) has the line legends underneath which I cannot collapse. Is there a way to a least collapse them? Thanks in advance.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How to show fit line graphs in a single report, without the tables and line legends
One additional method that can work, is to create a Journal Window (Shift/CNTL/J) and then copying just the parts of the display output you want and then pasting them into the new Journal Window
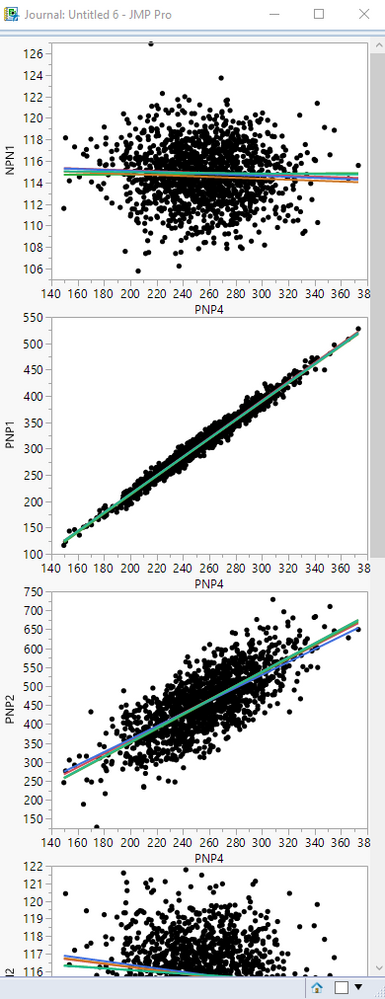
Below is the Journal Window the plots that were pasted into it
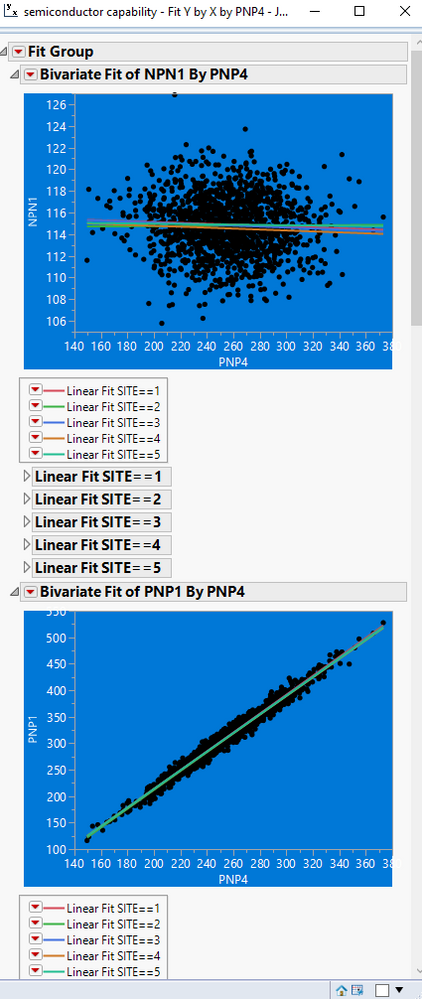
Below is a script that illustrates how to delete the items not wanted. With some work, this could be expanded on, and become a complete solution
Names Default To Here( 1 );
dt = Open( "$SAMPLE_DATA/Semiconductor Capability.jmp" );
biv = Bivariate(
Y( :NPN1 ),
X( :PNP1 ),
Fit Where( :SITE == 1, Fit Line( {Line Color( {213, 72, 87} )} ) ),
Fit Where( :SITE == 2, Fit Line( {Line Color( {57, 177, 67} )} ) ),
Fit Where( :SITE == 3, Fit Line( {Line Color( {64, 111, 223} )} ) ),
Fit Where( :SITE == 4, Fit Line( {Line Color( {207, 121, 38} )} ) ),
Fit Where( :SITE == 5, Fit Line( {Line Color( {33, 189, 145} )} ) ),
SendToReport(
Dispatch( {}, "Linear Fit SITE==1", OutlineBox, {Close( 1 )} ),
Dispatch( {}, "Linear Fit SITE==2", OutlineBox, {Close( 1 )} ),
Dispatch( {}, "Linear Fit SITE==3", OutlineBox, {Close( 1 )} ),
Dispatch( {}, "Linear Fit SITE==4", OutlineBox, {Close( 1 )} ),
Dispatch( {}, "Linear Fit SITE==5", OutlineBox, {Close( 1 )} )
)
);
Report( biv )[Picture Box( 2 )] << delete;
Report( biv )["Linear Fit Site==1"] << delete;
Report( biv )["Linear Fit Site==2"] << delete;
Report( biv )["Linear Fit Site==3"] << delete;
Report( biv )["Linear Fit Site==4"] << delete;
Report( biv )["Linear Fit Site==5"] << delete;
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: How to show fit line graphs in a single report, without the tables and line legends
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us