- New to JMP? Let the Data Analysis Director guide you through selecting an analysis task, an analysis goal, and a data type. Available now in the JMP Marketplace!
- See how to install JMP Marketplace extensions to customize and enhance JMP.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- Dose Response Probit analysis
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Dose Response Probit analysis
Hi group,
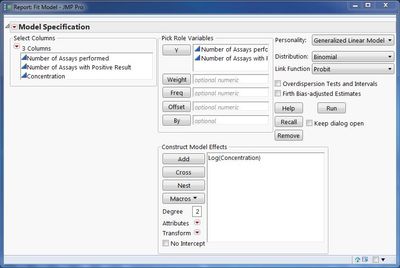
I am trying to reproduce a simple dose response analysis that I had performed in SPSS. So in JMP i am using the "Fit model" module, selecting for Generalized linear models, binomial distribution and probit link.
I have three variables, number of assays performed (10 assays), number of assays with positive result and concentration (from 0 to 36)
0 10 2
1 10 4
2 10 9
6 10 16
8 10 25
9 10 36
0 10 0
I select number of assays performed as "Y" variable, number of assays with positive result as "Freq" and concentration in logs as variable to model effects on. But when I hit run I get this message in the results:
"Warning: Hessian not positive definite. Singularity is likely due to a poor fit of the model, or linear dependencies among the model covariates. Fit and result are of questionable value"
And the analysis is not performed.
Can anyone share some comments?
Thanks,
Dave
Accepted Solutions
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Dose Response Probit analysis
Dave,
Your "number of assays with positive result" needs to go in as a "Y" variable also. The Frequency role is for indicating how many times a row in the data table actually occurred. In this case, the GLM model expects the number of assays performed and the number of positive results as "separate" Y values. The analysis will link them together.
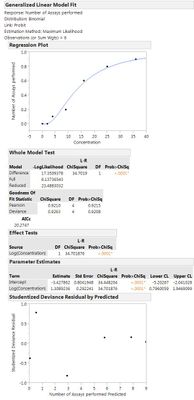
One thing you did not mention that is often done for this type of work is to take the log of the concentration. I did this in the model dialog shown above. Highlight the concentration model effect, click the transform button and choose log. Results are posted here.
Dan
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Dose Response Probit analysis
Dave,
Your "number of assays with positive result" needs to go in as a "Y" variable also. The Frequency role is for indicating how many times a row in the data table actually occurred. In this case, the GLM model expects the number of assays performed and the number of positive results as "separate" Y values. The analysis will link them together.
One thing you did not mention that is often done for this type of work is to take the log of the concentration. I did this in the model dialog shown above. Highlight the concentration model effect, click the transform button and choose log. Results are posted here.
Dan
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Dose Response Probit analysis
Thanks a lot DanO, that is a very clear answer. It is interesting that JMP knows they are linked.
Regarding the log transformation, yes I will take that into account (it was indeed mentioned in my original post, although pretty vague I must admit!)
Best,
Dave
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Dose Response Probit analysis
Dear DanO,
sorry for bothering you again, but changing the numbers to the demo, I seem to be getting wrong results. Please have a look at this dataset, with same variables but different numbers. For the three highest concentrations I get 100% positives
Number of positive results Number of assays performed Concentration
0 1140 0.01
241 1140 0.1
590 1140 0.5
963 1140 1
1134 1140 5
1140 1140 10
1140 1140 50
1140 1140 100
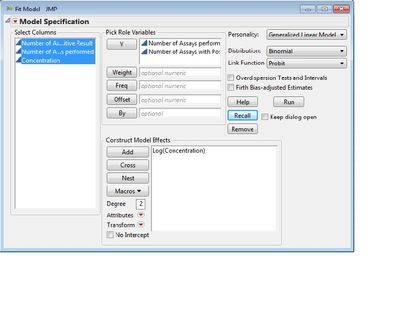
I run the Probit as you suggested
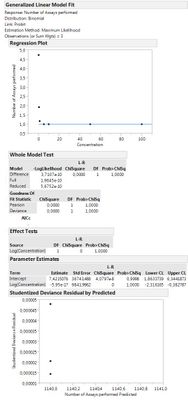
An after running I get a strange result which doesn't make sense to me. Instead of doing percentages of corrects on the Y, it seems to be doing something different. What am I doing wrong?
Thanks ever so much for your help
Dave
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Dose Response Probit analysis
Been following this thread with interest.
From the JMP docs:
If your data is summarized as frequencies of events and trials, specify two continuous
columns in this order: a count of the number of successes, and a count of the number of
trials. Alternatively, you can specify the number of failures instead of successes.
(my emphasis).
So if you switch the order the 2 y variables are entered I think you will see more of what you expected especially if you change the x axis to be logarithmic.
That said, I can't explain why the order doesn't seem to matter with your original data.
Michael
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Dose Response Probit analysis
The 0 10 0 last row from the original data for some reason insulates the analysis from needing the correct order. If you leave it out then order does matter.
Michael
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Dose Response Probit analysis
That is indeed the right order to feed in the variables. Thanks for pointing that out, Michael.
Dave
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us