- Learn how to build custom Python data connectors and further customize JMP’s Data Connector Framework with the Python Data Connector Demo, available now in the JMP Marketplace!
- See how to create experiments to support product design and ID useful product features. Register for June 12 webinar, 2pm US Eastern Time.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Mute
- Printer Friendly Page
Discussions
Solve problems, and share tips and tricks with other JMP users.- JMP User Community
- :
- Discussions
- :
- Analysis by specific rows
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Analysis by specific rows
Hi Everyone!
I have a very large dataset that I am in the midst of analyzing but I have come across some difficulties.
In the dataset there are 1924 rows and 1061 columns - so it's quite large. I am trying to do Fit Model and just want to make sure that I am doing this right.
If I for example want to fit a model based on rows 112-245, I select these rows, enter "Fit Model" option, select the column and right click on it, select "rows" -> "selected" (which will create a new column named "Selected") and perform the analysis, right?
And when I do this, I want to add some variables to the model in "Construct model effects". Will the variables that I select also automatically be for the rows 112-245 or do I need to do something else for this to happen?
Hope that it makes sense!
Cheers!
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
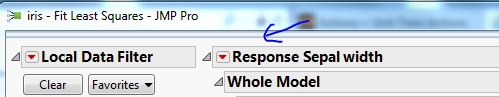
One more step, you still need to filter the rows being analyzed: Once the fit model platform comes up, hover over the bar at the top of the screen so the menu is visible, then add a local data filter. Choose your 'Selected' column and press Add, then click on the value '1'.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
Thanks for getting back to me!
I am not sure that I follow completely, though.
When I enter the Fit Model option, this is what I get:
I add the new "Selected" column that I have created as Y and then throw in the variables that I want to add (in this case sex, age etc) in the Contruct Model Effects box. I don't see any of the options that you you mention?
When I apply the analysis, I get the statistical results but nothing that looks similar to yours.
Cheers
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
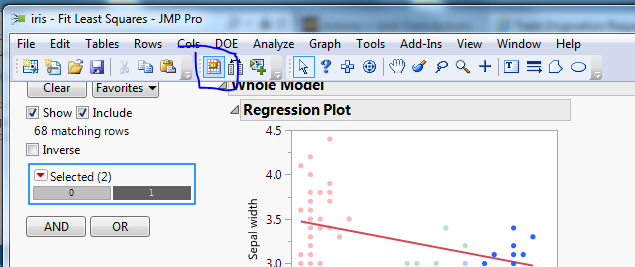
If you can't view the screenshot in my previous post, here it is again
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
Once you have created the column for the row selection, I'd suggest putting that column in the "By" box for Fit Model. You will get a seperate (but identical in terms of the model construction) analysis for the selected and unselected rows. This allows you to only focus on the selected rows model or to compare the models for the selected and unselected rows.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
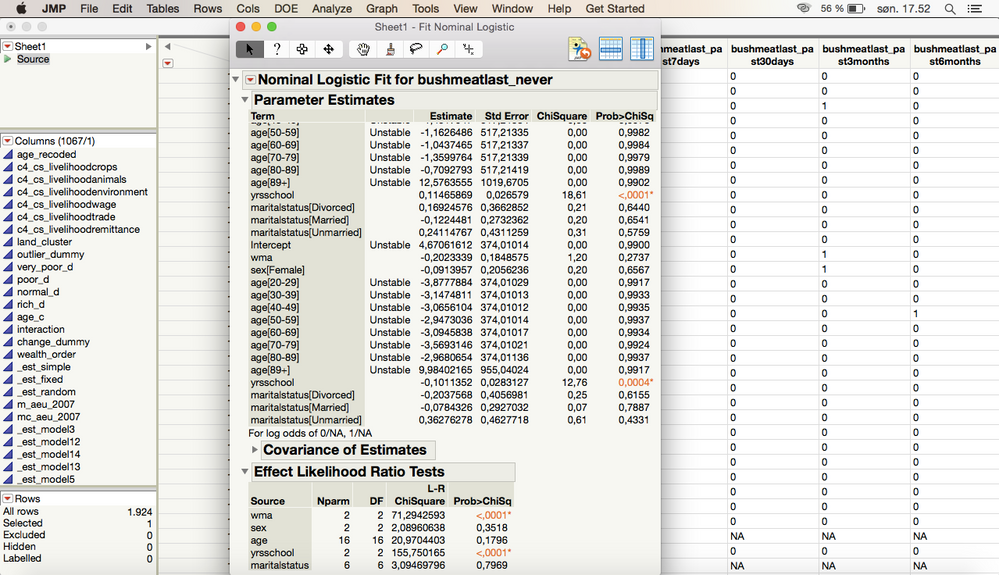
OK, so I ran a test on the column that I am interested in on specific rows that I had selected and got this:
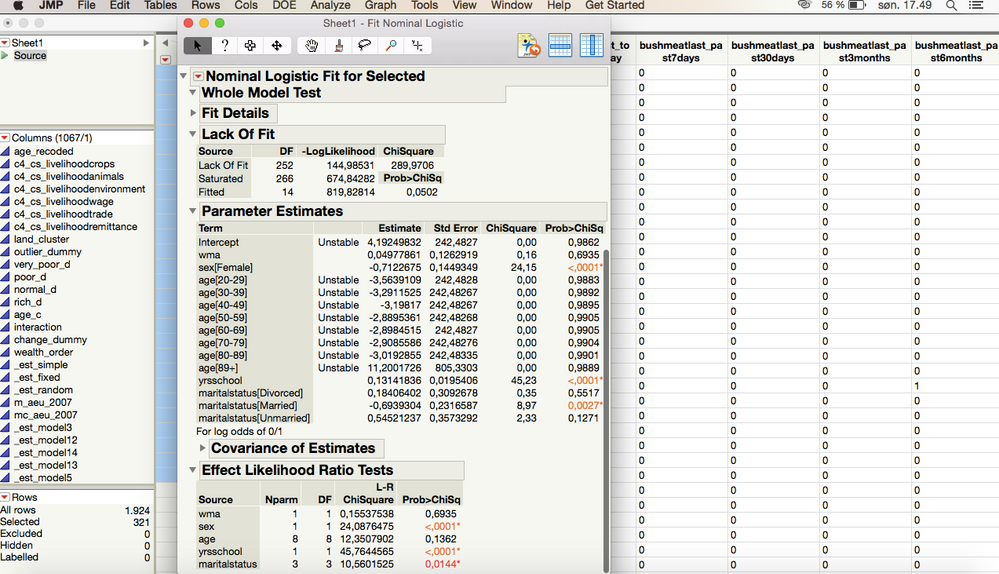
I also tried doing the model for the entire column, without selecting specific rows (which would be the simplest way for me to analyze my data) and it came out as this
For some reason when I do this, the model is analyzed twice in the Parameter Estimates, which I don't understand why?
Cheers
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
I don't know about the analysis appearing twice, but I see two other problems. Both of your analyses (the one for the selected rows and the one for the whole column) are using all of the rows. Once you select the rows, you must either subset them to do an analysis just on those rows - or, as I am suggesting, make the selected rows a new column (1=selected, 0=not selected) and put that column in the By box when you launch the platform. Then you will get two analyses - one for the selected rows and one for the others and you can compare them.
The second problem is that your model is trying to fit a response variable with 3 levels - 0, 1, and NA. While this can be done, I think you really are tryiing to build a model to predict the 1 values. Your variables appear twice because they are trying to predict 1/NA and 0/NA. It would help if you would post a screenshot of the Fit Model dialog you are using.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
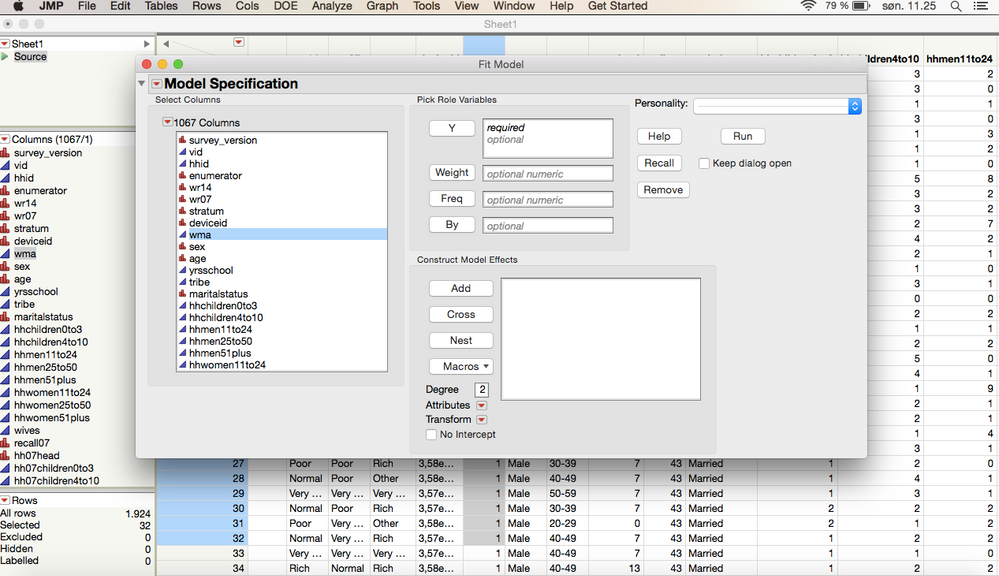
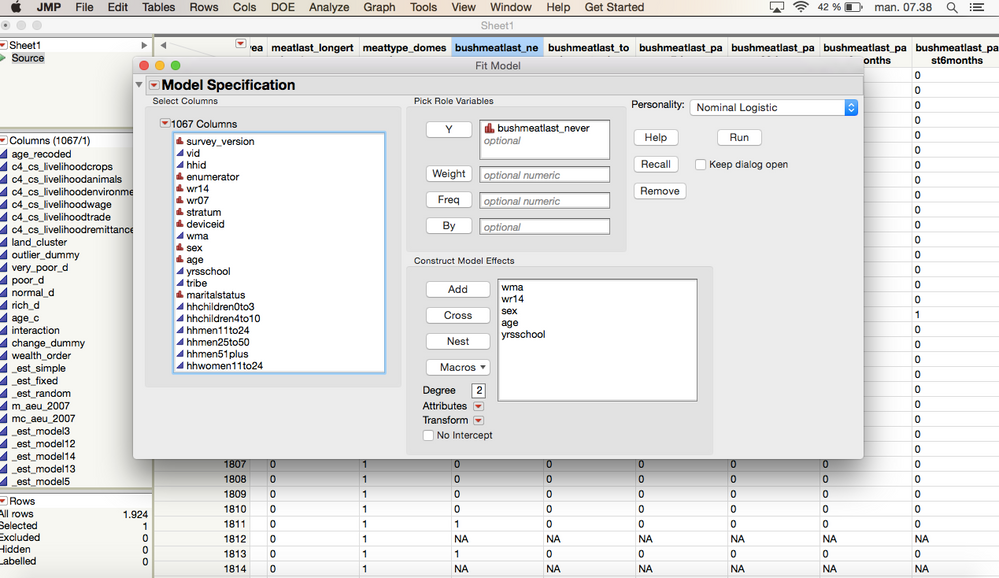
So this is how it would look before I run the test:
I want to test which factors affect the column (Bushmeat never consumed). The variables are wma(0 and 1), wr14 (very poor, poor, normal, rich), sex (male, female), age, yrsschool and marital status (Married, unmarried, widower, divorced).
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
How many levels does your Bushmeat column have? From what you have said, I think you want to, but your prior analysis makes me think there are 3 levels - if NA is in that column, you should change it to missing. Also, the box near the middle of the Fit Model Box says "By" and that is where you should put the column that identifies the selected rows. That is what will give you a separate analysis for the selected and unselected rows. If you don't care about that comparison and only want to analyze the selected rows, subset them before doing the Fit Model.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Get Direct Link
- Report Inappropriate Content
Re: Analysis by specific rows
The Bushmeat column has two levels, yes. 0 and 1 corresponding to a yes or no answer. The N/A is if there was no answer given, so I think you are right that I should remove those. I did that and that took care of the problem with the analysis being run twice.
Thanks!
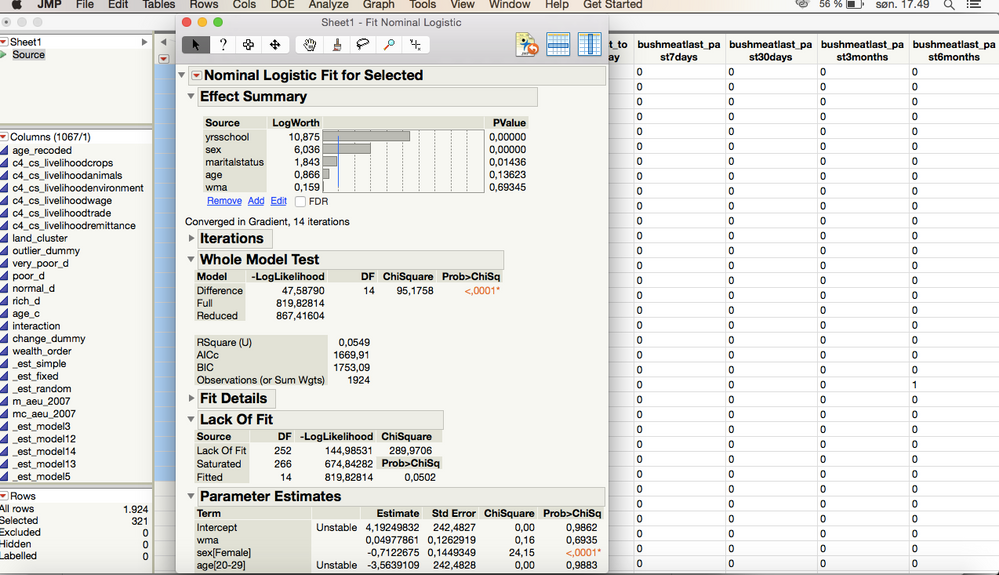
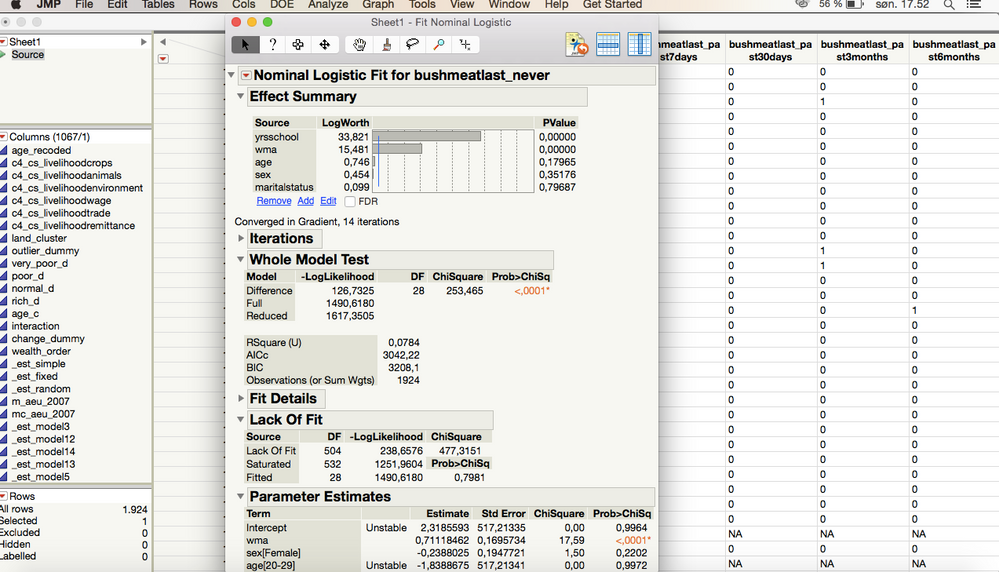
As of the analysis itself, I get this result for one of the tests
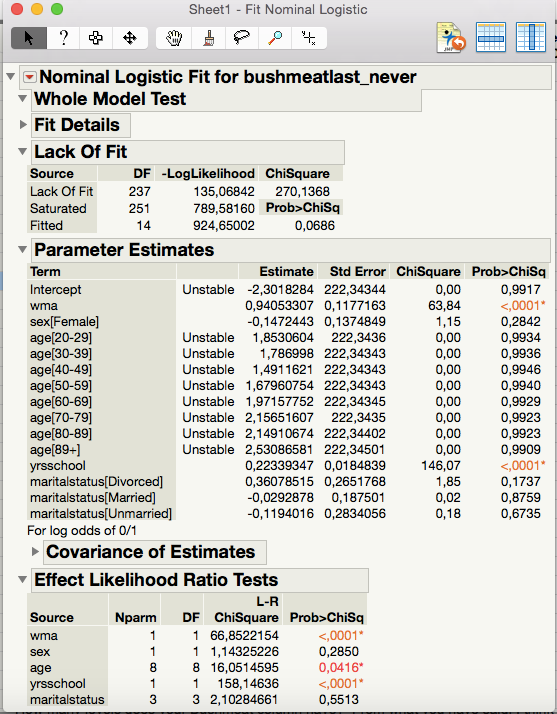
I interpert this as there being a significant effect of the variables "wma" and "yrsschool" and that the Effect Likelihood Ratio Tests indicates that for "wma" it is for the value being 1 that is significant, as well as 1 being the value of significance for "yrsschool". Correct?
Recommended Articles
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us