Turn on suggestions
Auto-suggest helps you quickly narrow down your search results by suggesting possible matches as you type.
Options
- Subscribe
- Mark as read
- Mark as new
Learn JMP Events
Events designed to further your knowledge and exploration of JMP.- JMP User Community
- :
- Learn JMP
- :
- Learn JMP Events
- :
- JMP Academic Webinar - Genomics Research with JMP Pro
JMP Academic Webinar - Genomics Research with JMP Pro
Presented in English

Get JMP software free for academic use at jmp.com/student
JMP Pro 17 marked a significant change in how genomic data analysis is performed in JMP. New platforms were introduced for genetic marker analysis and breeding simulations, and several existing platforms were optimized for the handling of large, wide data sets. Unlike the specialized JMP Genomics software, these new features are built directly into JMP Pro and do not require a SAS backend. This webinar demonstrates how to apply JMP Pro in genomics research through case studies on Differential Gene Expression and GWAS.
Start:
Tue, Apr 4, 2023 01:00 PM EDT
End:
Tue, Apr 4, 2023 02:00 PM EDT
2 Comments
02-21-2024
12:03 PM
02-21-2024
12:03 PM
Thanks for the amazing webinar Chris! I sincerely appreciate the work from the JMP Academic Team to provide resources to the academic community. I have been using the platform and your workflow to analyze a proteomics dataset in JMP17Pro and had the following questions:
- You used the hierarchical clustering platform with Hybrid Ward as a QC screen and to explore missing values. When I try to do this on my data set of ~1000 mitochondrial proteins, I receive an error that there are too many missing values to cluster and I am basically forced to impute values to do any clustering. Why did this error not show up for you when you clustered to identify missing values before removing columns with too much missing data?
- I have also been trying to modify the script in your journal file that removes columns with too many zeros (my data has blanks and not zeros but same concept, I think). I need a further level of evaluation as I have 13 rows of Control samples and 13 rows of Experimental samples in the data and want to filter (or create 2 additional new groupings like you did) for any columns of a protein in which “Control Missing Too Many Values” and “Experimental Missing Too Many Values”. Sometimes when a protein is predominantly missing from samples in one condition and present in another, that could be biologically interesting so I would not necessarily want that filtered because the column as a whole did not have 60% non-blanks. I know a little R for coding but am struggling to find a way to do this in JSL.
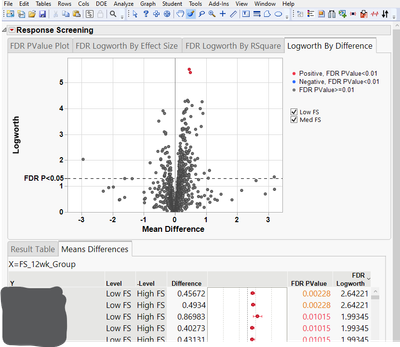
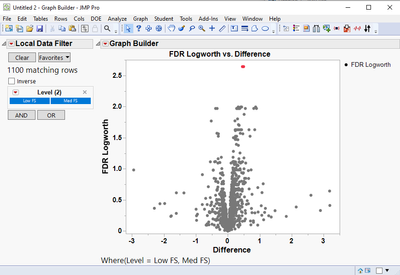
- Is it possible to change the graphing on the volcano plots in the Response Screening to color FDR Pvalues < 0.02 or <0.05 (and not the defaulting 0.01)? I can change the location of the reference line (see image below) but it would be nice to have points colored consistent with an FDR cutoff of choice and also have the legend updated on the Logworth by Difference plot. I thought a workaround would be to generate a new table of the differences (log2 FC) and P-values and take them into GraphBuilder where I have more control but I get the same coloring of only the points <0.01 so the coloring setting seems to follow the table (see image below).
Thank you for any assistance,
TK
02-22-2024
11:32 AM
02-22-2024
11:32 AM
Hi @TimboK1 ,
Thanks for watching the webinar. I will do my best to answer your questions.
- In my example for the cattle data, it did not have any missing data they were 0s. I misspoke by calling them missing as in true blank values. If they were true missing data then using Missing Data Pattern underneath the Table menu or Analyze>Screening> Explore Missing Values would be a better start. There are a lot of options for imputing and even clustering for missingness. In your case, you would have to use the Missing value imputation option or recode the missing values to 0 if they really 0 or not measured/present. This will cause some interesting differences and confidence in those differences. Sometimes the original data is a 0 and then normalization methods that have a log transform step will cause these values to be missing. In these cases an offset is usually used (by adding a 1 before transforming) so the final transformed value is still 0 as opposed to a real missing value (issue with measuring the sample).
- Yes, I agree that one has to segment the groups in these cases. I prefer to do the search by group given that the amount of missing or 0's can be disproportionate to a particular group. A by group option would be good to add, but would require a little extra coding. I can take a look at what it would take to do this in JSL, but it will take me a little time as to the best approach (and to put a little user interface option for a By Group column).
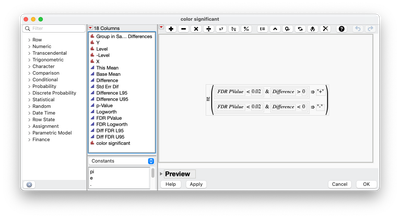
- For JMP Pro 17 or JMP Pro 18 (released in March), this is currently not possible. I do something similar as you did. I save out the Means difference table (Red Triangle within Response Screening and choose Save Tables>Save Means Differences) and then use graph builder with a local data filter. I also will use Rows>Row Selection>Select Where to select rows that are below a threshold of choice and difference threshold. Then I use Rows>Row Selection>Name Selection to Column to create an indicator column(s) that I can use to concatenate for a column I can use in the color option in graph builder. Another option is to use a column formula where the conditions are added and an if statement. I like this option better since it is a single action and saved within the column as to what I am doing.
Hope that helps.
Chris Kirchberg, M.S.2
Data Scientist, Life Sciences - Global Technical Enablement
JMP Statistical Discovery, LLC. - Denver, CO
Tel: +1-919-531-9927 ▪ Mobile: +1-303-378-7419 ▪ E-mail: chris.kirchberg@jmp.com
www.jmp.com
Data Scientist, Life Sciences - Global Technical Enablement
JMP Statistical Discovery, LLC. - Denver, CO
Tel: +1-919-531-9927 ▪ Mobile: +1-303-378-7419 ▪ E-mail: chris.kirchberg@jmp.com
www.jmp.com
- © 2026 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- Contact Us