Hi @PriorChipmunk23 ,
Here is one way that might help you:
In Fit Curve you can find the closest model (as you already did) and save out the parametric prediction formula:
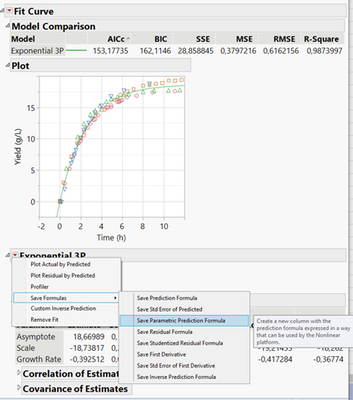
You can then go to the Nonlinear Platform, using this formula as the predictor, and your output variable as Y. Hit “Go” to start the fitting process, and then you can save prediction formula, Standard Error, formulae for confidence limits or just the limit values:
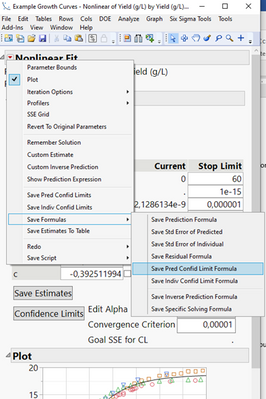
If you want to do this by batch, simply put the column with the batch-ID in the “By”-Window in the launch dialogue. It will then launch as many fitting dialogues as you have levels in your Batch-ID column.
Hope this helps!
Florian