The QTL Single Marker Analysis tool in JMP Genomics performs a simple regression to correlate quantitative traits with markers within a genome. This tool is a great way to scan an entire set of markers and quickly identify loci of interest for a trait. In this example we will analyze 90 recombinant inbred lines of oat for their mean differential scanning calorimetry (DSC). Note that the two data sets used as input for this analysis, as well as the linkage groups defined in those sets, were created in the post Genetic Association with JMP Genomics, Part 4: Basic Linkage Mapping.
- From the Genomics Starter menu, select Genetics > Linkage Maps and QTL > QTL Single Marker Analysis.
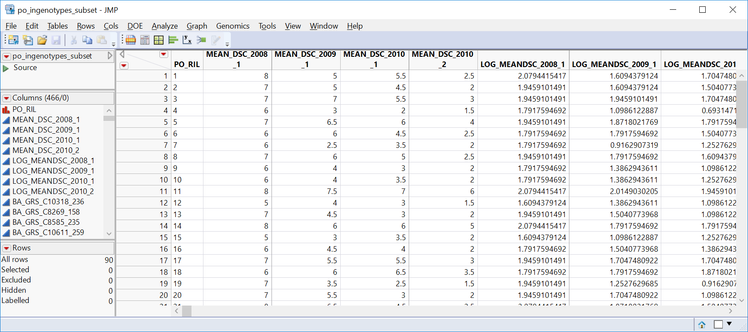
- In the General tab, Choose po_ingenotypes_subst.sas7bdat as the Input SAS Data Set.
- This is a wide data set containing 4 mean DSC measurements and ~450 markers from 90 oat lines.
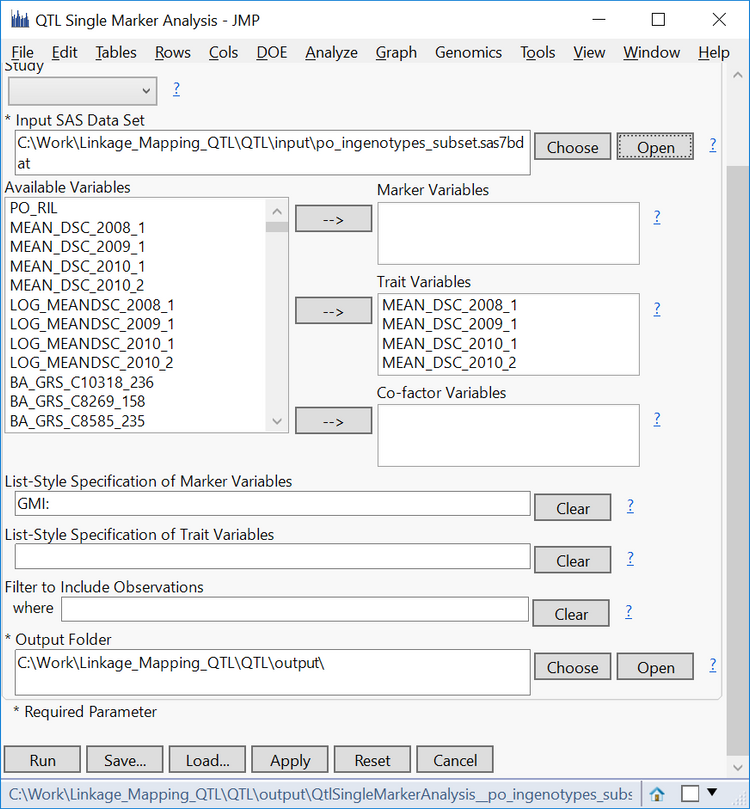
- From the Available Variables box, select the four “MEAN_DSC” variables and move them into the Trait Variables
- Enter “GMI:” into the List-Style Specification of Marker Variables field to designate every column beginning with “GMI” as a column containing marker data.
- Specify an Output Folder.
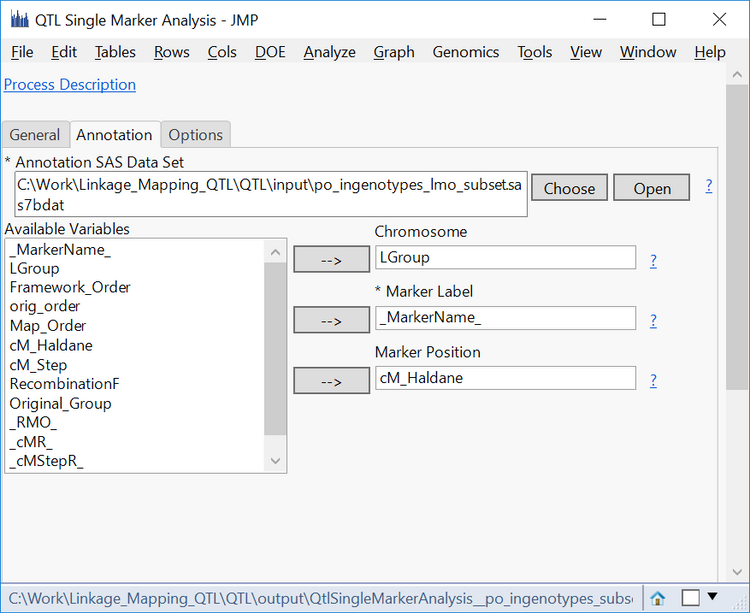
- In the Annotation tab, choose po_ingenotypes_lmo_subset.sas7bdat as the Annotation SAS Data Set.
- Select LGroup as the Chromosome, _MarkerName_ as the Marker Label, and cM_Haldane as the Marker Position.
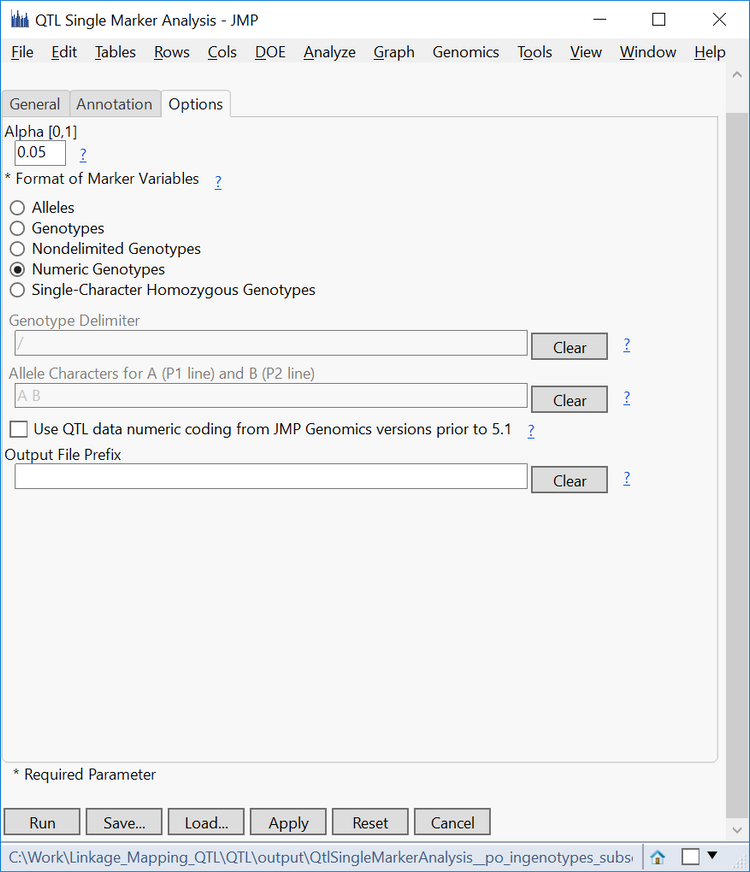
- In the Options tab, set a standard Alpha level of 05.
- Select Numeric Genotypes as the Format of Marker Variables.
- Click Run at the bottom of the window to begin the analysis.
Results
- The initial output window displays four histograms. One for each of the four DSC means measured in the input data set. These histograms show the number of significant markers in each linkage group for the trait specified.
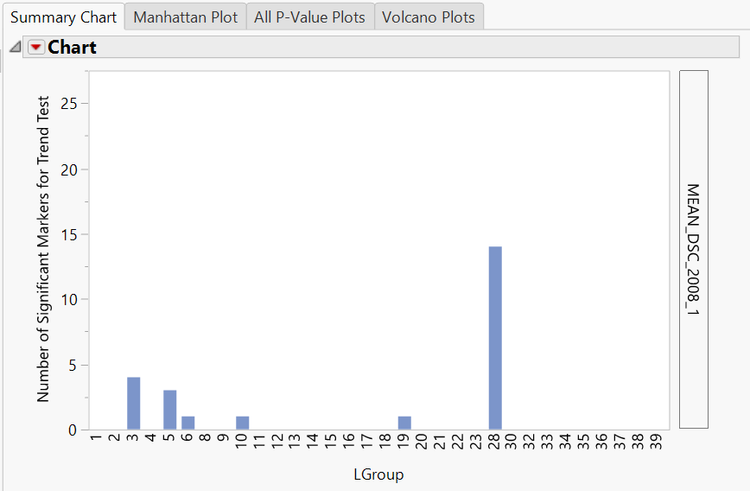
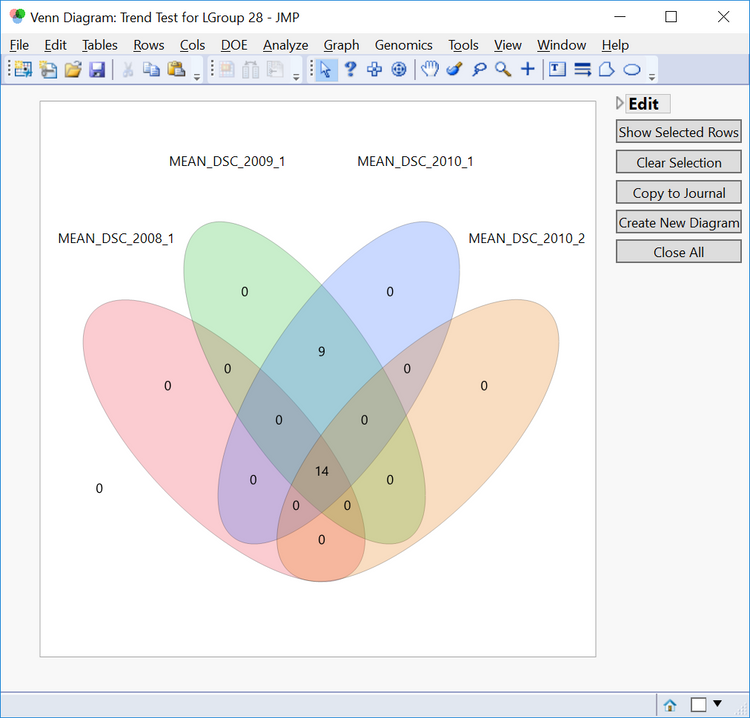
- Linkage group 28 appears to have several significant markers for each trait. In order to inspect these markers further, click the Trend button under View Venn Diagram of Significant Markers in the Drill Downs. A venn diagram will appear showing the 25 markers and which traits they are significant for. To create a subset of these markers, click any of the non-zero numbers in the venn diagram and select Show Selected Rows.
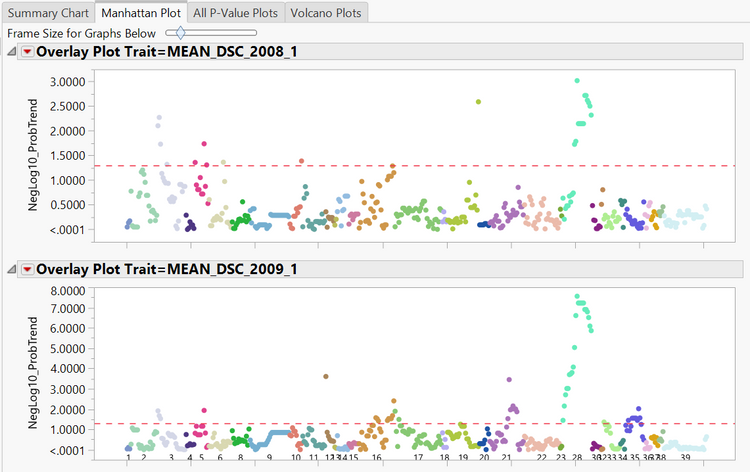
- In the Manhattan Plot tab, a manhattan plot is given for each trait with significant markers for each linkage group being shown above the dotted red line (representing the 0.05 alpha level). Hover over the points on each plot to see the marker name.
- The All P-Value Plots tab gives a plot of individual markers in each linkage group for each trait.
- The Volcano Plots tab shows the significance of each marker with significant markers for each linkage group being shown above the dotted red line and points to the left of the plots representing increased incidence of the minor allele and points to the right of the plots representing increased incidence of the major allele. Selecting any linkage group or groups from the LGroup menu on the right will highlight only points from those groups in the plots.
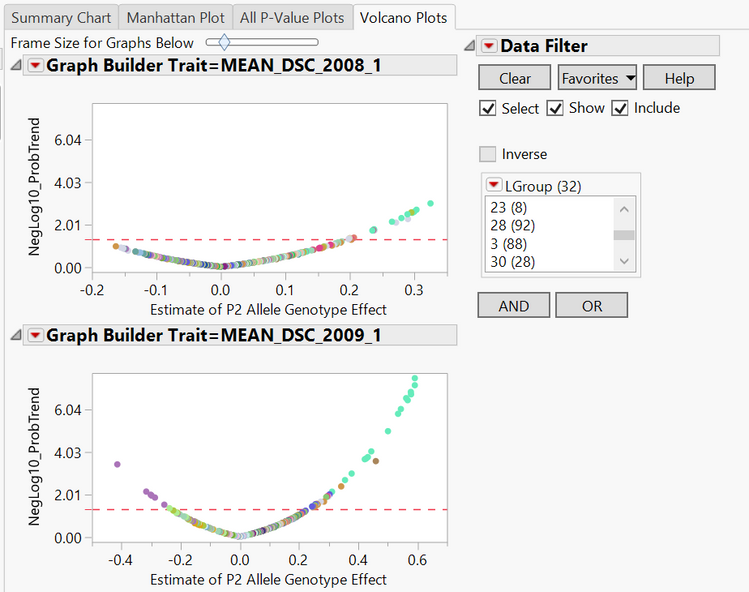
*The interactive reports generated from this workflow can be viewed on JMP Public.
Summary
The QTL Single Marker Analysis tool is a great way to perform a quick scan of an entire set of marker data for QTL signals. The software does a simple regression for each marker with values for each trait. In order to carry out the analysis, a genotype data set is needed with marker information and phenotypic information in wide format, as well as an annotation data set with linkage group and position info. Subsets of markers of interest can be easily created using the Drill Downs menu.
You must be a registered user to add a comment. If you've already registered, sign in. Otherwise, register and sign in.