- JMP User Community
- :
- JMP Discovery Summit Series
- :
- Past Discovery Summits
- :
- Discovery Summit Europe 2018 Presentations
- :
- Model Validation Strategies for Designed Experiments Using Bootstrapping Techniq...
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
Level: Intermediate
Philip Ramsey, Principal Lecturer, University of New Hampshire; and Owner, North Haven Group
Chris Gotwalt, JMP Director of Statistical Research and Development, SAS
There are two different goals to statistical modeling: explanation and prediction. Explanatory models often predict poorly (Shmueli, 2010). Often analyses of designed experiments (DOE) are explanatory, yet the experimental goals are prediction. DOE is a best practice for product and process development where one predicts future performance. Predictive modeling requires partitioning the data into training and validation sets where the validation set is used to assess predictive models. Most DOEs have insufficient observations to form a validation set precluding direct assessment of prediction performance. We demonstrate a “balanced auto-validation” technique using the original data to create two copies of that data, one a training set and the other a validation set. The sets differ in row weightings. The weights are Dirichlet distributed and “balanced;” observations contributing more to the training data contribute less to the validation set (and vice versa). The approach only requires copying the data and creating a formula column for weights. The technique is general, allowing one to apply predictive modeling techniques to smaller data sets common to laboratory and manufacturing studies. Two biopharma process development case studies are used for demonstration. Both cases have large validation sets combined with definitive screening designs. JMP is used to demonstrate analyses.
This was a really interesting presentation. Thanks for sharing the files. I am trying to reproduce this on one of my DOEs.
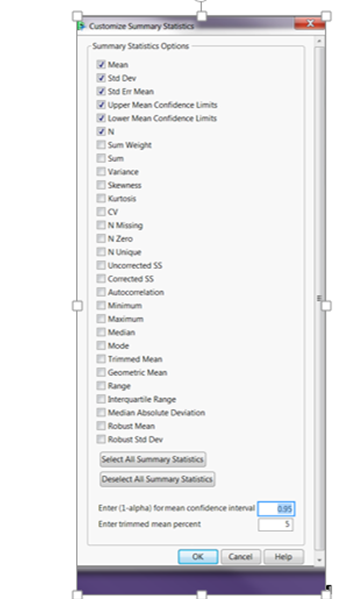
However I got stuck at step 10 from the slides: Calculate the proportion non zero. That option is not available for me (see picture below)?? Does anyone know why that might be the case? I am using JMP Pro 14EA.
I have got around this by using N Zero instead and making an extra calculation to get the proportion non zero. Works nicely thanks!!
Yes, the easy solution is to use the NZero function (number of times 0)
and then you can calculate the proportion zero or nonzero. If you look
at the case studies, then you will see I used the NZero function to
create various weighting schemes.
Phil Ramsey
Thanks Phil. I have now downloaded JMP14, and the option for proportion on zero is now included :)
Thanks Phil and Chris - This has been fun to play with for my DOE data. Is there any reason not to do this with DOE data plus historical data? The sample sets are still small and would not make good designs if analyzed in DOE platform - could I just make duplicate sets of the data I have and follow all steps? or is there a reason not to do that - Thanks Patti
Really nice to hear you are having fun with the autovalidation method. The method is very general and can easily be applied to observational or DOE data and I encourage you to try. The method is done exactly the same way whether observational or DOE. I also have applied it to small observational datasets and have gotten very useful results in terms of finding which effects appear most important.
Phil Ramsey
I am having so much fun with it – it is distracting me from trying out the Functional Data Explorer – which I want to try next. It takes a lot of time to run the model simulations – but I think it is well worth it for what I am doing and helping us identify important factors for further experimentation!
Thank you!
Patti
Really pleased that you are finding autovalidation useful and fun .
Phil
this technique appears to be a game-changer for medical researchers who struggle with small sample issues. are there any citations of articles published in peer-reviewed journals that I can be referred to? many thanks for this gift.
- © 2024 JMP Statistical Discovery LLC. All Rights Reserved.
- Terms of Use
- Privacy Statement
- About JMP
- JMP Software
- JMP User Community
- Contact